Note
Go to the end to download the full example code. or to run this example in your browser via Binder
Using brain templates from TemplateFlow¶
In this example, we see how to fetch and use brain templates from TemplateFlow.
We first see how to fetch and use the Desikan-Killiany (DK) atlas in order to make surface plots.
Then, we see how to get the proper template for the Harvard-Oxford volumetric atlas.
Datasets used in this example
Fetch DK Surface template¶
TemplateFlow allows for the retrieval of image files, annotations, and
metadata for various brain templates.
Specific files are accessed via the get function
by providing the template name
and filtering by file name ‘entities’
such as resolution (res),
suffix, atlas, hemisphere (hemi), space,
description (desc)…
Tips
TemplateFlow files follow a BIDS filename template and should more less be organized as BIDS template.
If a search matches multiple files, the function returns a list of all matching paths.
Available resources can be explored in the TemplateFlow browser.
import templateflow.api as tflow
from nilearn.surface import SurfaceImage, load_surf_data
template = "fsaverage"
fetched_files = tflow.get(
template,
extension="surf.gii",
suffix="pial",
)
print(fetched_files)
Downloading https://templateflow.s3.amazonaws.com/tpl-fsaverage/tpl-fsaverage_hemi-L_den-3k_pial.surf.gii
0%| | 0.00/66.1k [00:00<?, ?B/s]
100%|██████████| 66.1k/66.1k [00:00<00:00, 1.02MB/s]
Downloading https://templateflow.s3.amazonaws.com/tpl-fsaverage/tpl-fsaverage_hemi-L_den-10k_pial.surf.gii
0%| | 0.00/271k [00:00<?, ?B/s]
19%|█▉ | 52.2k/271k [00:00<00:00, 399kB/s]
65%|██████▌ | 176k/271k [00:00<00:00, 718kB/s]
100%|██████████| 271k/271k [00:00<00:00, 823kB/s]
Downloading https://templateflow.s3.amazonaws.com/tpl-fsaverage/tpl-fsaverage_hemi-L_den-41k_pial.surf.gii
0%| | 0.00/1.07M [00:00<?, ?B/s]
7%|▋ | 71.7k/1.07M [00:00<00:01, 513kB/s]
20%|█▉ | 211k/1.07M [00:00<00:01, 796kB/s]
41%|████ | 437k/1.07M [00:00<00:00, 1.17MB/s]
75%|███████▌ | 803k/1.07M [00:00<00:00, 1.74MB/s]
100%|██████████| 1.07M/1.07M [00:00<00:00, 1.90MB/s]
Downloading https://templateflow.s3.amazonaws.com/tpl-fsaverage/tpl-fsaverage_hemi-L_den-164k_pial.surf.gii
0%| | 0.00/4.34M [00:00<?, ?B/s]
2%|▏ | 69.6k/4.34M [00:00<00:08, 505kB/s]
5%|▍ | 209k/4.34M [00:00<00:05, 796kB/s]
12%|█▏ | 505k/4.34M [00:00<00:02, 1.40MB/s]
22%|██▏ | 975k/4.34M [00:00<00:01, 2.18MB/s]
38%|███▊ | 1.64M/4.34M [00:00<00:00, 3.10MB/s]
60%|█████▉ | 2.59M/4.34M [00:00<00:00, 4.37MB/s]
100%|██████████| 4.34M/4.34M [00:00<00:00, 4.75MB/s]
Downloading https://templateflow.s3.amazonaws.com/tpl-fsaverage/tpl-fsaverage_hemi-R_den-3k_pial.surf.gii
0%| | 0.00/66.1k [00:00<?, ?B/s]
100%|██████████| 66.1k/66.1k [00:00<00:00, 1.02MB/s]
Downloading https://templateflow.s3.amazonaws.com/tpl-fsaverage/tpl-fsaverage_hemi-R_den-10k_pial.surf.gii
0%| | 0.00/271k [00:00<?, ?B/s]
26%|██▌ | 69.6k/271k [00:00<00:00, 518kB/s]
64%|██████▍ | 174k/271k [00:00<00:00, 667kB/s]
100%|██████████| 271k/271k [00:00<00:00, 999kB/s]
Downloading https://templateflow.s3.amazonaws.com/tpl-fsaverage/tpl-fsaverage_hemi-R_den-41k_pial.surf.gii
0%| | 0.00/1.07M [00:00<?, ?B/s]
7%|▋ | 69.6k/1.07M [00:00<00:01, 501kB/s]
18%|█▊ | 191k/1.07M [00:00<00:01, 719kB/s]
39%|███▉ | 418k/1.07M [00:00<00:00, 1.13MB/s]
80%|███████▉ | 853k/1.07M [00:00<00:00, 1.91MB/s]
100%|██████████| 1.07M/1.07M [00:00<00:00, 1.90MB/s]
Downloading https://templateflow.s3.amazonaws.com/tpl-fsaverage/tpl-fsaverage_hemi-R_den-164k_pial.surf.gii
0%| | 0.00/4.34M [00:00<?, ?B/s]
2%|▏ | 69.6k/4.34M [00:00<00:08, 530kB/s]
5%|▍ | 209k/4.34M [00:00<00:04, 944kB/s]
10%|█ | 452k/4.34M [00:00<00:02, 1.38MB/s]
21%|██ | 905k/4.34M [00:00<00:01, 2.22MB/s]
40%|████ | 1.74M/4.34M [00:00<00:00, 3.73MB/s]
70%|██████▉ | 3.04M/4.34M [00:00<00:00, 5.82MB/s]
100%|██████████| 4.34M/4.34M [00:00<00:00, 5.04MB/s]
[PosixPath('/home/runner/.cache/templateflow/tpl-fsaverage/tpl-fsaverage_hemi-L_den-3k_pial.surf.gii'), PosixPath('/home/runner/.cache/templateflow/tpl-fsaverage/tpl-fsaverage_hemi-L_den-10k_pial.surf.gii'), PosixPath('/home/runner/.cache/templateflow/tpl-fsaverage/tpl-fsaverage_hemi-L_den-41k_pial.surf.gii'), PosixPath('/home/runner/.cache/templateflow/tpl-fsaverage/tpl-fsaverage_hemi-L_den-164k_pial.surf.gii'), PosixPath('/home/runner/.cache/templateflow/tpl-fsaverage/tpl-fsaverage_hemi-R_den-3k_pial.surf.gii'), PosixPath('/home/runner/.cache/templateflow/tpl-fsaverage/tpl-fsaverage_hemi-R_den-10k_pial.surf.gii'), PosixPath('/home/runner/.cache/templateflow/tpl-fsaverage/tpl-fsaverage_hemi-R_den-41k_pial.surf.gii'), PosixPath('/home/runner/.cache/templateflow/tpl-fsaverage/tpl-fsaverage_hemi-R_den-164k_pial.surf.gii')]
In this example, the Desikan-Killiany (DK) atlas is fetched. DK is defined with reference to the FreeSurfer average (fsaverage) template, which is provided at multiple densities (number of vertices per hemisphere): 3k, 10k, 41k, 164k Here, we build a SurfaceImage containing the labels for both hemispheres for the 3k and 10k densities.
used_densities = ["3k", "10k"]
# uncomment the following line in case you want to see all densities
# used_densities = ["3k", "10k", "41k", "164k"]
desikan_dict = {}
for density in ["3k", "10k"]:
mesh = {}
data = {}
for hemi in ["left", "right"]:
mesh[hemi] = tflow.get(
template,
extension="surf.gii",
suffix="pial",
density=density,
hemi=hemi[0].upper(),
)
desc = "curated" if density == "164k" else None
roi_data = load_surf_data(
tflow.get(
template,
atlas="Desikan2006",
density=density,
hemi=hemi[0].upper(),
extension="label.gii",
desc=desc,
)
)
if density == "164k":
roi_data[roi_data < 0] = 0
data[hemi] = roi_data
desikan_dict[density] = SurfaceImage(mesh=mesh, data=data)
Downloading https://templateflow.s3.amazonaws.com/tpl-fsaverage/tpl-fsaverage_hemi-L_den-3k_atlas-Desikan2006_seg-aparc_dseg.label.gii
0%| | 0.00/6.79k [00:00<?, ?B/s]
100%|██████████| 6.79k/6.79k [00:00<00:00, 18.9MB/s]
Downloading https://templateflow.s3.amazonaws.com/tpl-fsaverage/tpl-fsaverage_hemi-R_den-3k_atlas-Desikan2006_seg-aparc_dseg.label.gii
0%| | 0.00/6.79k [00:00<?, ?B/s]
100%|██████████| 6.79k/6.79k [00:00<00:00, 26.7MB/s]
Downloading https://templateflow.s3.amazonaws.com/tpl-fsaverage/tpl-fsaverage_hemi-L_den-10k_atlas-Desikan2006_seg-aparc_dseg.label.gii
0%| | 0.00/9.61k [00:00<?, ?B/s]
100%|██████████| 9.61k/9.61k [00:00<00:00, 34.2MB/s]
Downloading https://templateflow.s3.amazonaws.com/tpl-fsaverage/tpl-fsaverage_hemi-R_den-10k_atlas-Desikan2006_seg-aparc_dseg.label.gii
0%| | 0.00/9.52k [00:00<?, ?B/s]
100%|██████████| 9.52k/9.52k [00:00<00:00, 27.5MB/s]
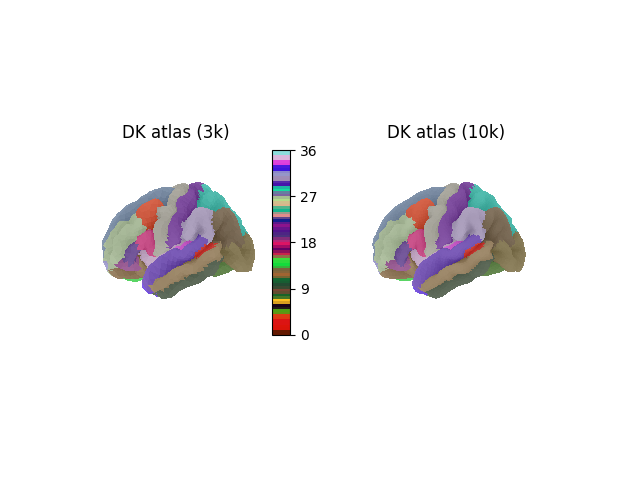
Plotting the DK atlas¶
To plot the DK atlas, we also fetch the lookup table (LUT) from TemplateFlow, to map region indices to the original region names and colors. We also use the background sulcal depth map to add more detail to the surface plot. To do this we map the density to the corresponding fsaverage data that Nilearn can fetch directly.
import matplotlib.pyplot as plt
from nilearn.datasets import load_fsaverage_data
from nilearn.plotting import plot_surf_roi, show
_, axes = plt.subplots(1, 2, subplot_kw={"projection": "3d"})
fs_density = {
"3k": "fsaverage4",
"10k": "fsaverage5",
"41k": "fsaverage6",
"164k": "fsaverage",
}
lut = tflow.get(
template,
atlas="Desikan2006",
suffix="dseg",
extension="tsv",
)
for ax, density in zip(axes, ["3k", "10k"], strict=True):
desikan = desikan_dict[density]
sulcal_depth_map = load_fsaverage_data(
mesh=fs_density[density], data_type="sulcal"
)
plot_surf_roi(
roi_map=desikan,
cmap=lut,
bg_map=sulcal_depth_map,
bg_on_data=True,
title=f"DK atlas ({density})",
axes=ax,
)
if density == "10k":
ax._colorbars[0].remove()
show()
[load_fsaverage_data] Dataset created in /home/runner/nilearn_data/fsaverage4
[load_fsaverage_data] Downloading data from https://osf.io/gkqd5/download ...
[load_fsaverage_data] ...done. (3 seconds, 0 min)
[load_fsaverage_data] Extracting data from /home/runner/nilearn_data/fsaverage4/
70c52c9f064022a30aee9395723e66be/download...
[load_fsaverage_data] .. done.
[load_fsaverage_data] Dataset directory found:
/home/runner/nilearn_data/fsaverage4
/home/runner/work/nilearn/nilearn/examples/01_plotting/plot_templateflow.py:141: RuntimeWarning:
Meshes are not identical but have compatible number of vertices.
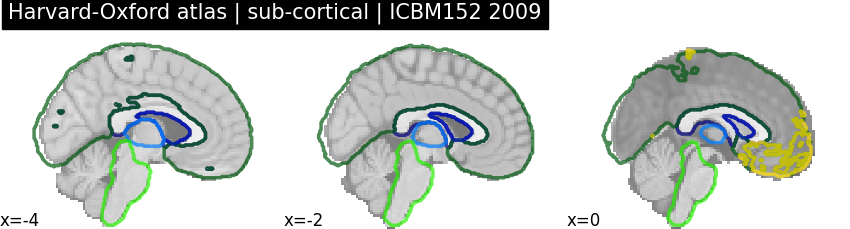
Getting the right template for volumetric atlases¶
The template used by nilearn to plot volumetric images is ICBM152 2009, release a. However, some atlases are defined on different templates, this is the case for the Harvard-Oxford atlas. This can lead to some imprecisions when plotting volumetric atlases, with regions encompassing non-brain areas.
from nilearn.datasets import fetch_atlas_harvard_oxford
from nilearn.plotting import plot_roi
plotting_params = {
"view_type": "contours",
"display_mode": "x",
"cut_coords": [-4, -2, 0],
"black_bg": False,
}
harvard_oxford_sub = fetch_atlas_harvard_oxford("sub-maxprob-thr25-1mm")
plot_roi(
harvard_oxford_sub.filename,
title="Harvard-Oxford atlas | sub-cortical | ICBM152 2009",
**plotting_params,
)
[fetch_atlas_harvard_oxford] Dataset directory found:
/home/runner/nilearn_data/fsl
<nilearn.plotting.displays._slicers.XSlicer object at 0x7f69f5c063e0>
This is because the template of the Harvard-Oxford atlas is:
print(f"Harvard-Oxford atlas template: {harvard_oxford_sub.template}")
Harvard-Oxford atlas template: MNI152NLin6Asym
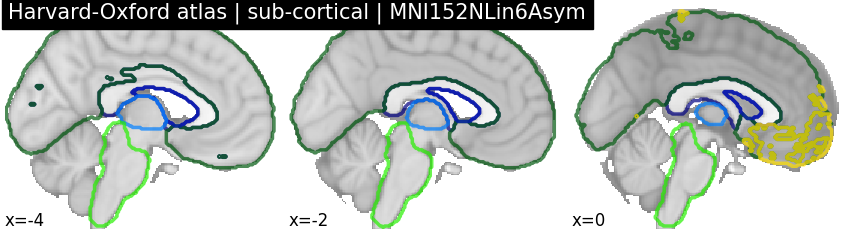
Getting a template¶
If you want to visualize the Harvard-Oxford atlas on the template it was originally defined, you can get it from TemplateFlow. This template is the default template used by FSL, and its unique identifier within TemplateFlow is`MNI152NLin6Asym`.
template = "MNI152NLin6Asym"
resolution = "01"
MNI152NLin6Asym_template_img = tflow.get(
template,
resolution=resolution,
suffix="T1w",
desc="brain", # Set desc-brain to get the skull-stripped variant
extension="nii.gz",
)
print(f"{MNI152NLin6Asym_template_img=}")
plot_roi(
harvard_oxford_sub.filename,
title="Harvard-Oxford atlas | sub-cortical | MNI152NLin6Asym",
bg_img=MNI152NLin6Asym_template_img,
**plotting_params,
)
show()
Downloading https://templateflow.s3.amazonaws.com/tpl-MNI152NLin6Asym/tpl-MNI152NLin6Asym_res-01_desc-brain_T1w.nii.gz
0%| | 0.00/3.29M [00:00<?, ?B/s]
2%|▏ | 69.6k/3.29M [00:00<00:05, 537kB/s]
7%|▋ | 244k/3.29M [00:00<00:03, 1.01MB/s]
20%|██ | 662k/3.29M [00:00<00:01, 2.01MB/s]
40%|████ | 1.32M/3.29M [00:00<00:00, 3.21MB/s]
71%|███████ | 2.34M/3.29M [00:00<00:00, 4.87MB/s]
100%|██████████| 3.29M/3.29M [00:00<00:00, 4.57MB/s]
MNI152NLin6Asym_template_img=PosixPath('/home/runner/.cache/templateflow/tpl-MNI152NLin6Asym/tpl-MNI152NLin6Asym_res-01_desc-brain_T1w.nii.gz')
Total running time of the script: (0 minutes 28.400 seconds)
Estimated memory usage: 165 MB