Note
Go to the end to download the full example code. or to run this example in your browser via Binder
Predicted time series and residuals¶
Here we fit a First Level GLM with the minimize_memory-argument set to False. By doing so, the FirstLevelModel-object stores the residuals, which we can then inspect. Also, the predicted time series can be extracted, which is useful to assess the quality of the model fit.
Import modules¶
import pandas as pd
from nilearn import image, masking
from nilearn.datasets import fetch_spm_auditory
from nilearn.plotting import plot_stat_map, show
# load fMRI data
subject_data = fetch_spm_auditory()
fmri_img = subject_data.func[0]
# Make an average
mean_img = image.mean_img(fmri_img)
mask = masking.compute_epi_mask(mean_img)
# Clean and smooth data
fmri_img = image.clean_img(fmri_img, standardize=None)
fmri_img = image.smooth_img(fmri_img, 5.0)
# load events
events = pd.read_csv(subject_data.events, sep="\t")
[fetch_spm_auditory] Dataset directory found:
/home/runner/nilearn_data/spm_auditory
Fit model¶
Note that minimize_memory is set to False so that FirstLevelModel stores the residuals. signal_scaling is set to False, so we keep the same scaling as the original data in fmri_img.
from nilearn.glm.first_level import FirstLevelModel
fmri_glm = FirstLevelModel(
t_r=subject_data.t_r,
drift_model="cosine",
signal_scaling=False,
mask_img=mask,
minimize_memory=False,
verbose=1,
)
fmri_glm = fmri_glm.fit(fmri_img, events)
/home/runner/work/nilearn/nilearn/examples/04_glm_first_level/plot_predictions_residuals.py:57: RuntimeWarning:
[MultiNiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[FirstLevelModel.fit] Computing run 1 out of 1 runs (go take a coffee, a big
one).
\[FirstLevelModel.fit] Performing mask computation.
\[FirstLevelModel.fit] Masking took 0 seconds.
\[FirstLevelModel.fit] Performing GLM computation.
\[FirstLevelModel.fit] GLM took 1 seconds.
\[FirstLevelModel.fit] Computation of 1 runs done in 00 HR 00 MIN 01 SEC.
Calculate and plot contrast¶
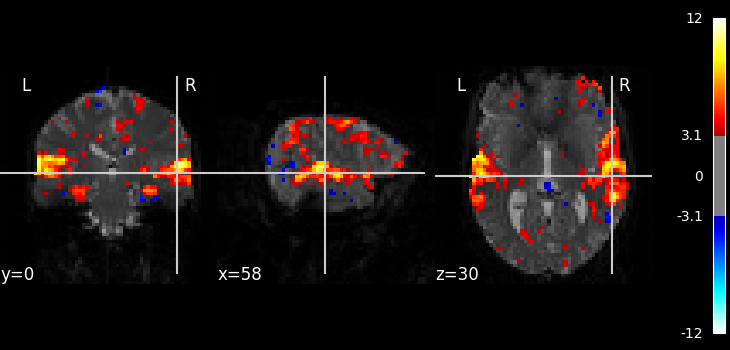
z_map = fmri_glm.compute_contrast("listening")
threshold = 3.1
plot_stat_map(
z_map,
bg_img=mean_img,
threshold=threshold,
title=f"listening > rest (t-test; |Z|>{threshold})",
)
show()
Extract the largest clusters¶
We can extract the 6 largest clusters surviving our threshold. and get the x, y, and z coordinates of their peaks. We then extract the time series from a sphere around each coordinate.
from nilearn.maskers import NiftiSpheresMasker
from nilearn.reporting import get_clusters_table
table = get_clusters_table(
z_map, stat_threshold=threshold, cluster_threshold=20
)
table.set_index("Cluster ID", drop=True)
print(table)
coords = table.loc[range(1, 7), ["X", "Y", "Z"]].to_numpy()
print(coords)
masker = NiftiSpheresMasker(coords, verbose=1)
real_timeseries = masker.fit_transform(fmri_img)
predicted_timeseries = masker.fit_transform(fmri_glm.predicted[0])
Cluster ID X Y Z Peak Stat Cluster Size (mm3)
0 1 -60.0 -6.0 42.0 11.543910 20196
1 1a -51.0 -12.0 39.0 10.497958
2 1b -39.0 -12.0 39.0 10.118978
3 1c -63.0 12.0 33.0 9.754835
4 2 60.0 0.0 36.0 11.088600 32157
.. ... ... ... ... ... ...
58 21 27.0 -72.0 27.0 4.422543 729
59 21a 24.0 -66.0 18.0 4.346647
60 21b 24.0 -78.0 21.0 3.428392
61 22 -30.0 -24.0 90.0 3.877083 648
62 22a -33.0 -18.0 72.0 3.791863
[63 rows x 6 columns]
[[-51. -12. 39.]
[-39. -12. 39.]
[-63. 12. 33.]
[ 60. 0. 36.]
[ 66. 12. 27.]
[ 60. -18. 30.]]
\[NiftiSpheresMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/04_glm_first_level/plot_predictions_residuals.py:96: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
\[NiftiSpheresMasker.wrapped] Loading data from <nibabel.nifti1.Nifti1Image
object at 0x7f5bc0a844c0>
\[NiftiSpheresMasker.wrapped] Extracting region signals
\[NiftiSpheresMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/04_glm_first_level/plot_predictions_residuals.py:96: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
/home/runner/work/nilearn/nilearn/.tox/doc/lib/python3.10/site-packages/nibabel/onetime.py:156: FutureWarning:
residuals' is deprecated.
It will be removed in Nilearn 0.16.0.
Use 'residuals_' instead.
\[NiftiSpheresMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/04_glm_first_level/plot_predictions_residuals.py:97: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
\[NiftiSpheresMasker.wrapped] Loading data from <nibabel.nifti1.Nifti1Image
object at 0x7f5be27d96c0>
\[NiftiSpheresMasker.wrapped] Extracting region signals
\[NiftiSpheresMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/04_glm_first_level/plot_predictions_residuals.py:97: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
Let’s have a look at the report to make sure the spheres are well placed.
/home/runner/work/nilearn/nilearn/examples/04_glm_first_level/plot_predictions_residuals.py:102: UserWarning:
`generate_report()` received 'displayed_spheres=10' to be displayed. But masker only has 6 sphere(s). 'displayed_spheres' was set to 6.
Plot predicted and actual time series for 6 most significant clusters¶
import matplotlib.pyplot as plt
# colors for each of the clusters
colors = ["blue", "navy", "purple", "magenta", "olive", "teal"]
# plot the time series and corresponding locations
fig1, axs1 = plt.subplots(2, 6)
for i in range(6):
# plotting time series
axs1[0, i].set_title(f"Cluster peak {coords[i]}\n")
axs1[0, i].plot(real_timeseries[:, i], c=colors[i], lw=2)
axs1[0, i].plot(predicted_timeseries[:, i], c="r", ls="--", lw=2)
axs1[0, i].set_xlabel("Time")
axs1[0, i].set_ylabel("Signal intensity", labelpad=0)
# plotting image below the time series
roi_img = plot_stat_map(
z_map,
cut_coords=[coords[i][2]],
threshold=3.1,
figure=fig1,
axes=axs1[1, i],
display_mode="z",
colorbar=False,
bg_img=mean_img,
)
roi_img.add_markers([coords[i]], colors[i], 300)
fig1.set_size_inches(24, 14)
show()
![Cluster peak [-51. -12. 39.] , Cluster peak [-39. -12. 39.] , Cluster peak [-63. 12. 33.] , Cluster peak [60. 0. 36.] , Cluster peak [66. 12. 27.] , Cluster peak [ 60. -18. 30.]](../../_images/sphx_glr_plot_predictions_residuals_002.png)
Get residuals¶
\[NiftiSpheresMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/04_glm_first_level/plot_predictions_residuals.py:142: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
\[NiftiSpheresMasker.wrapped] Loading data from <nibabel.nifti1.Nifti1Image
object at 0x7f5be3d991b0>
\[NiftiSpheresMasker.wrapped] Extracting region signals
\[NiftiSpheresMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/04_glm_first_level/plot_predictions_residuals.py:142: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
Plot distribution of residuals¶
Note that residuals are not really distributed normally.
![Cluster peak [-51. -12. 39.] , Cluster peak [-39. -12. 39.] , Cluster peak [-63. 12. 33.] , Cluster peak [60. 0. 36.] , Cluster peak [66. 12. 27.] , Cluster peak [ 60. -18. 30.]](../../_images/sphx_glr_plot_predictions_residuals_003.png)
Mean residuals: 0.018869405610299532
Mean residuals: 0.04575602612203456
Mean residuals: -1.554312234475219e-14
Mean residuals: -0.13042393047259077
Mean residuals: -0.014998994819316611
Mean residuals: 0.04955626049962536
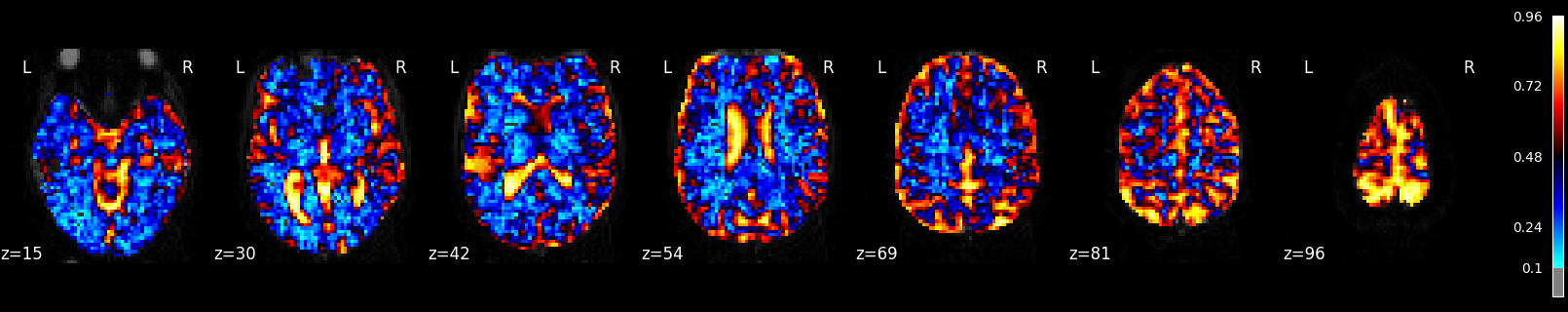
Plot R-squared¶
Because we stored the residuals, we can plot the R-squared: the proportion of explained variance of the GLM as a whole. Note that the R-squared is markedly lower deep down the brain, where there is more physiological noise and we are further away from the receive coils. However, R-Squared should be interpreted with a grain of salt. The R-squared value will necessarily increase with the addition of more factors (such as rest, active, drift, motion) into the GLM. Additionally, we are looking at the overall fit of the model, so we are unable to say whether a voxel/region has a large R-squared value because the voxel/region is responsive to the experiment (such as active or rest) or because the voxel/region fits the noise factors (such as drift or motion) that could be present in the GLM. To isolate the influence of the experiment, we can use an F-test as shown in the next section.
plot_stat_map(
fmri_glm.r_square[0],
bg_img=mean_img,
threshold=0.1,
display_mode="z",
cut_coords=7,
cmap="inferno",
title="R-squared",
vmin=0,
symmetric_cbar=False,
)
/home/runner/work/nilearn/nilearn/.tox/doc/lib/python3.10/site-packages/nibabel/onetime.py:156: FutureWarning:
residuals' is deprecated.
It will be removed in Nilearn 0.16.0.
Use 'residuals_' instead.
<nilearn.plotting.displays._slicers.ZSlicer object at 0x7f5bf5754d90>
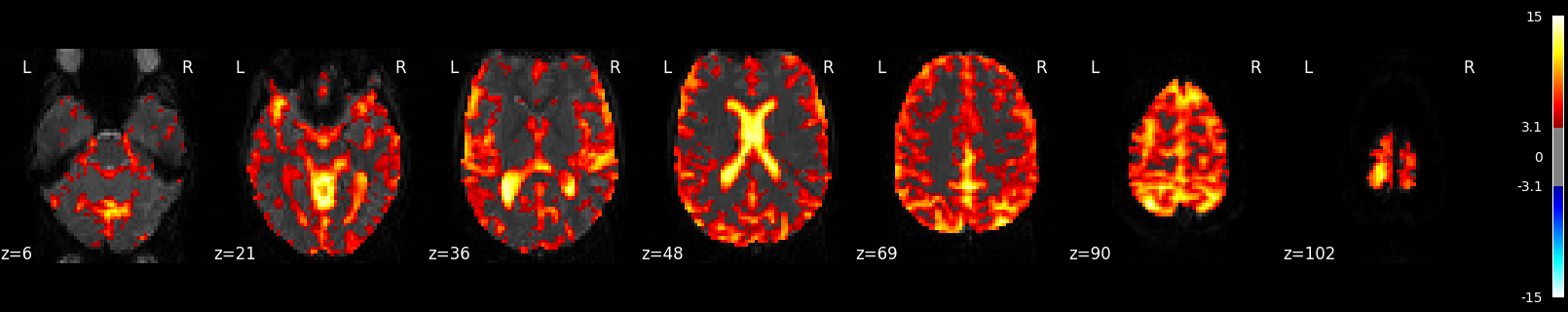
Calculate and Plot F-test¶
The F-test tells you how well the GLM fits effects of interest such as the active and rest conditions together. This is different from R-squared, which tells you how well the overall GLM fits the data, including active, rest and all the other columns in the design matrix such as drift and motion.
# f-test for 'listening'
z_map_ftest = fmri_glm.compute_contrast(
"listening", stat_type="F", output_type="z_score"
)
plot_stat_map(
z_map_ftest,
bg_img=mean_img,
threshold=threshold,
display_mode="z",
cut_coords=7,
cmap="inferno",
title=f"listening > rest (F-test; Z>{threshold})",
symmetric_cbar=False,
vmin=0,
)
show()
Total running time of the script: (0 minutes 17.826 seconds)
Estimated memory usage: 782 MB