Note
This page is a reference documentation. It only explains the function signature, and not how to use it. Please refer to the user guide for the big picture.
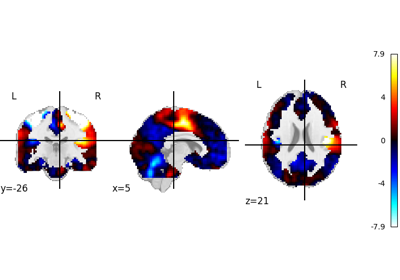
nilearn.plotting.plot_stat_map¶
- nilearn.plotting.plot_stat_map(stat_map_img, bg_img=<MNI152Template>, cut_coords=None, output_file=None, display_mode='ortho', colorbar=True, cbar_tick_format='%.2g', figure=None, axes=None, title=None, threshold=1e-06, annotate=True, draw_cross=True, black_bg='auto', cmap='RdBu_r', symmetric_cbar='auto', dim='auto', vmin=None, vmax=None, radiological=False, resampling_interpolation='continuous', transparency=None, transparency_range=None, **kwargs)[source]¶
Plot cuts of an ROI/mask image.
By default 3 cuts: Frontal, Axial, and Lateral.
- Parameters:
- stat_map_imgNiimg-like object
See Input and output: neuroimaging data representation. The statistical map image
- bg_imgNiimg-like object, optional
See Input and output: neuroimaging data representation. The background image to plot on top of. If nothing is specified, the MNI152 template will be used. To turn off background image, just pass “bg_img=None”. default=MNI152TEMPLATE.
- cut_coordsNone, allowed types depend on the
display_mode, optional The world coordinates of the point where the cut is performed.
If
display_modeis'ortho'or'tiled', this must be a 3-sequence offloatorint:(x, y, z).If
display_modeis'xz','yz'or'yx', this must be a 2-sequence offloatorint:(x, z),(y, z)or(x, y).If
display_modeis"x","y", or"z", this can be:If
display_modeis'mosaic', this can be:an
intin which case it specifies the number of cuts to perform in each direction"x","y","z".a 3-sequence of
floatorintin which case it specifies the number of cuts to perform in each direction"x","y","z"separately.dict<str: 1Dndarray> in which case keys are the directions (‘x’, ‘y’, ‘z’) and the values are sequences holding the cut coordinates.
If
Noneis given, the cuts are calculated automatically.
- output_file
strorpathlib.Pathor None, default=None The name of an image file to export the plot to. Valid extensions are .png, .pdf, .svg. If output_file is not None, the plot is saved to a file, and the display is closed.
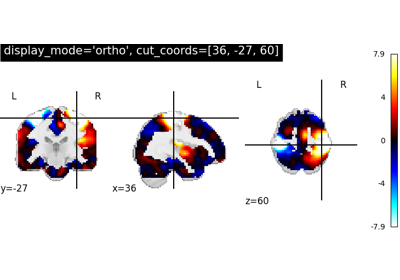
- display_mode{“ortho”, “tiled”, “mosaic”, “x”, “y”, “z”, “yx”, “xz”, “yz”}, default=”ortho”
Choose the direction of the cuts:
"x": sagittal"y": coronal"z": axial"ortho": three cuts are performed in orthogonal directions"tiled": three cuts are performed and arranged in a 2x2 grid"mosaic": three cuts are performed along multiple rows and columns
- colorbar
bool, optional If True, display a colorbar next to the plots. default=True.
- cbar_tick_format
str, default=”%.2g” (scientific notation) Controls how to format the tick labels of the colorbar. Ex: use “%i” to display as integers.
- figure
int, ormatplotlib.figure.Figure, or None, optional Matplotlib figure used or its number. If None is given, a new figure is created.
- axes
matplotlib.axes.Axes, or 4tupleoffloat: (xmin, ymin, width, height), default=None The axes, or the coordinates, in matplotlib figure space, of the axes used to display the plot. If None, the complete figure is used.
- title
str, or None, default=None The title displayed on the figure.
- threshold
intorfloat, None, or ‘auto’, optional If None is given, the image is not thresholded. If number is given, it must be non-negative. The specified value is used to threshold the image: values below the threshold (in absolute value) are plotted as transparent. If a string percentile is given, it should finish with percent sign e.g., “95%”. We threshold based on the score obtained using this percentile on the image data. If “auto” is given, the threshold is determined based on the score obtained using percentile value “80%” on the absolute value of the image data. default=1e-6.
- annotate
bool, default=True If annotate is True (like positions and / or left/right annotation) are added to the plot.
- draw_cross
bool, default=True If draw_cross is True, a cross is drawn on the plot to indicate the cut position.
- black_bg
bool, or “auto”, optional If True, the background of the image is set to be black. If you wish to save figures with a black background, you will need to pass facecolor=”k”, edgecolor=”k” to
matplotlib.pyplot.savefig. default=’auto’.- cmap
matplotlib.colors.Colormap, orstr, optional The colormap to use. Either a string which is a name of a matplotlib colormap, or a matplotlib colormap object.
default=default=”RdBu_r”.
- symmetric_cbar
bool, or “auto”, default=”auto” Specifies whether the colorbar and colormap should range from -vmax to vmax (or from vmin to -vmin if -vmin is greater than vmax) or from vmin to vmax. Setting to “auto” (the default) will select the former if either vmin or vmax is None and the image has both positive and negative values.
- dim
float, or “auto”, optional Dimming factor applied to background image. By default, automatic heuristics are applied based upon the background image intensity. Accepted float values, where a typical span is between -2 and 2 (-2 = increase contrast; 2 = decrease contrast), but larger values can be used for a more pronounced effect. 0 means no dimming. default=’auto’.
- vmin
floator obj:int or None, optional Lower bound of the colormap. The values below vmin are masked. If None, the min of the image is used. Passed to
matplotlib.pyplot.imshow.- vmax
floator obj:int or None, optional Upper bound of the colormap. The values above vmax are masked. If None, the max of the image is used. Passed to
matplotlib.pyplot.imshow.- resampling_interpolation
str, optional Interpolation to use when resampling the image to the destination space. Can be:
"continuous": use 3rd-order spline interpolation"nearest": use nearest-neighbor mapping.
Note
"nearest"is faster but can be noisier in some cases.default=’continuous’.
- radiological
bool, default=False Invert x axis and R L labels to plot sections as a radiological view. If False (default), the left hemisphere is on the left of a coronal image. If True, left hemisphere is on the right.
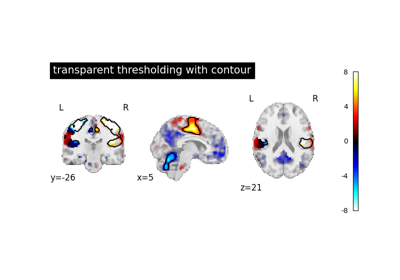
- transparency
floatbetween 0 and 1, or a Niimg-Like object, or None, default = None Value to be passed as alpha value to
imshow. ifNoneis passed, it will be set to 1. If an image is passed, voxel-wise alpha blending will be applied, by relying on the absolute value oftransparencyat each voxel.Added in Nilearn 0.12.0.
- transparency_range
tupleorlistof 2 non-negative numbers, or None, default = None When an image is passed to
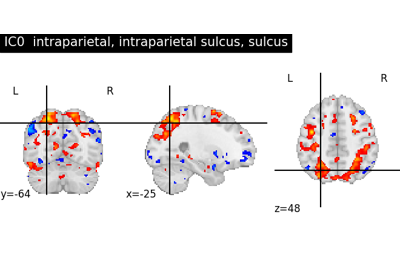
transparency, this determines the range of values in the image to use for transparency (alpha blending). For example withtransparency_range = [1.96, 3], any voxel / vertex (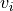 ):
):with a value between between -1.96 and 1.96, would be fully transparent (alpha = 0),
with a value less than -3 or greater than 3, would be fully opaque (alpha = 1),
with a value in the intervals
[-3.0, -1.96]or[1.96, 3.0], would have an alpha_i value scaled linearly between 0 and 1 :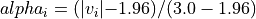 .
.
This parameter will be ignored unless an image is passed as
transparency. The first number must be greater than 0 and less than the second one. ifNoneis passed, this will be set to[0, max(abs(transparency))].Added in Nilearn 0.12.0.
- kwargsextra keyword arguments, optional
Extra keyword arguments ultimately passed to matplotlib.pyplot.imshow via
add_overlay.
- Returns:
- display
OrthoSliceror None An instance of the OrthoSlicer class. If
output_fileis defined, None is returned.
- display
- Raises:
- ValueError
if the specified threshold is a negative number
See also
nilearn.plotting.plot_anatTo simply plot anatomical images
nilearn.plotting.plot_epiTo simply plot raw EPI images
nilearn.plotting.plot_glass_brainTo plot maps in a glass brain
Notes
Arrays should be passed in numpy convention: (x, y, z) ordered.
For visualization, non-finite values found in passed ‘stat_map_img’ or ‘bg_img’ are set to zero.
Examples using nilearn.plotting.plot_stat_map¶
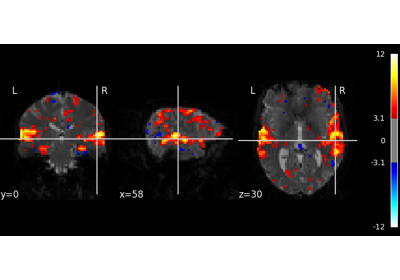
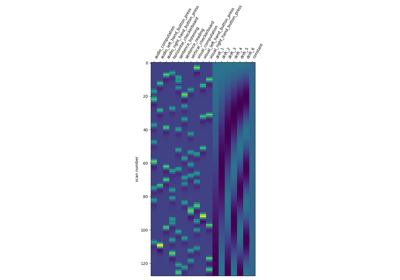
Intro to GLM Analysis: a single-run, single-subject fMRI dataset
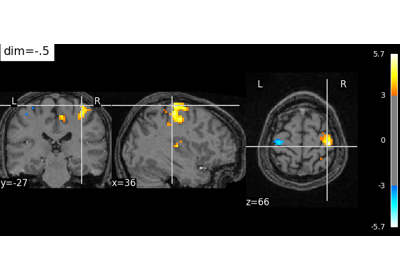
Controlling the contrast of the background when plotting
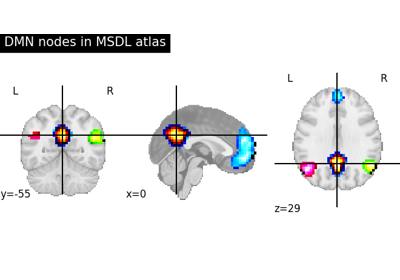
Visualizing a probabilistic atlas: the default mode in the MSDL atlas
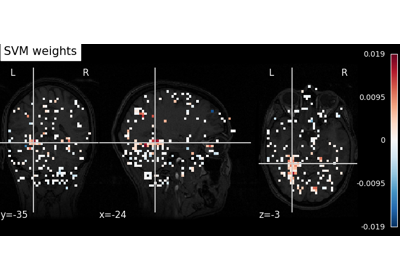
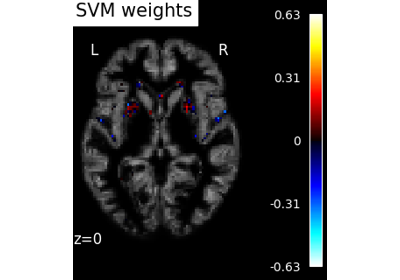
Decoding with ANOVA + SVM: face vs house in the Haxby dataset
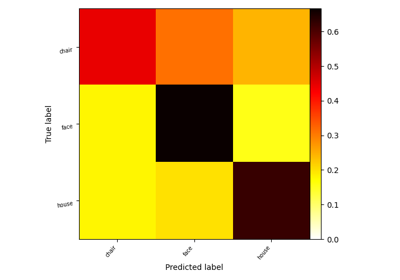
Decoding with FREM: face vs house vs chair object recognition
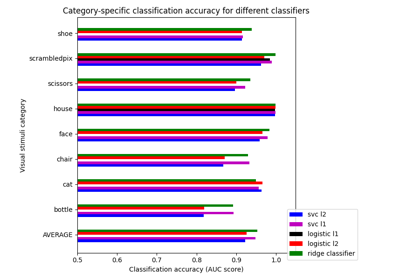
Different classifiers in decoding the Haxby dataset

Encoding models for visual stimuli from Miyawaki et al. 2008
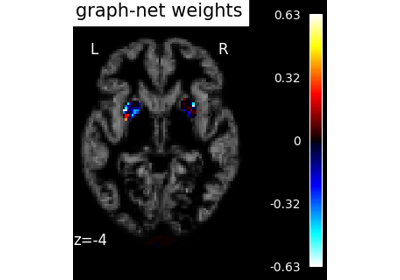
Voxel-Based Morphometry on Oasis dataset with Space-Net prior
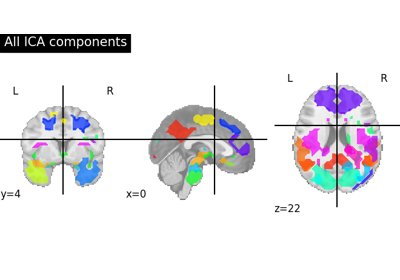
Deriving spatial maps from group fMRI data using ICA and Dictionary Learning
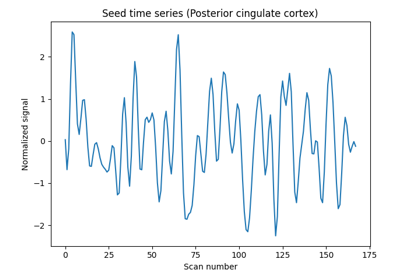
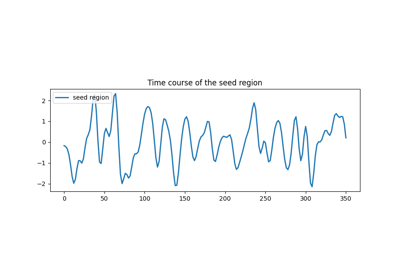
Producing single subject maps of seed-to-voxel correlation
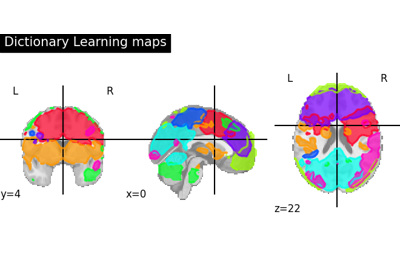
Regions extraction using dictionary learning and functional connectomes
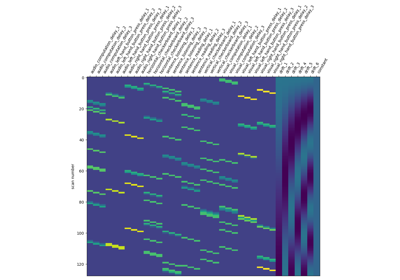
Analysis of an fMRI dataset with a Finite Impule Response (FIR) model
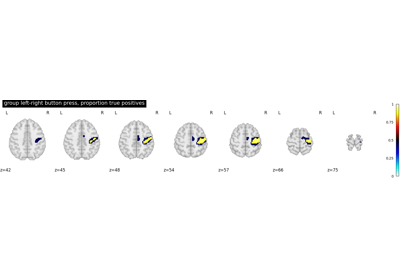
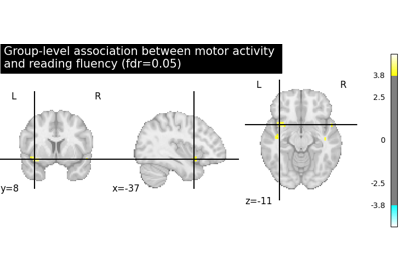
Second-level fMRI model: true positive proportion in clusters
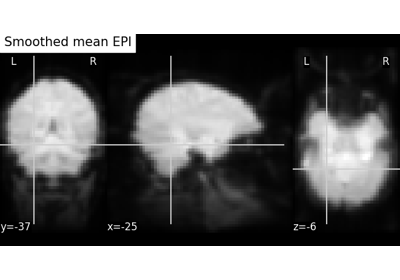
Computing a Region of Interest (ROI) mask manually
Regions Extraction of Default Mode Networks using Smith Atlas
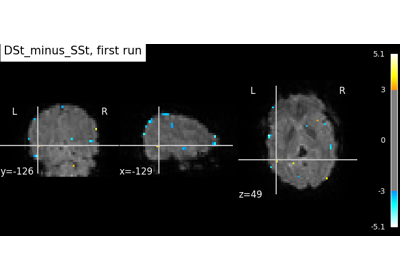
Beta-Series Modeling for Task-Based Functional Connectivity and Decoding
Massively univariate analysis of a calculation task from the Localizer dataset
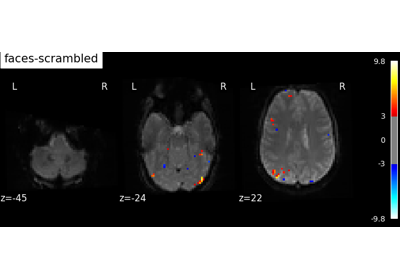
Massively univariate analysis of face vs house recognition
Multivariate decompositions: Independent component analysis of fMRI
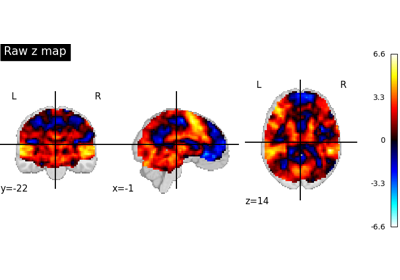
NeuroVault meta-analysis of stop-go paradigm studies