Note
Go to the end to download the full example code. or to run this example in your browser via Binder
Single-subject data (two runs) in native space¶
The example shows the analysis of an SPM dataset, with two conditions: viewing a face image or a scrambled face image.
This example takes a lot of time because the input are lists of 3D images sampled in different positions (encoded by different affine functions).
See also
For more information see the dataset description.
Fetch and inspect the data¶
Fetch the SPM multimodal_faces data.
from nilearn.datasets import fetch_spm_multimodal_fmri
subject_data = fetch_spm_multimodal_fmri()
[fetch_spm_multimodal_fmri] Dataset created in
/home/runner/nilearn_data/spm_multimodal_fmri
[fetch_spm_multimodal_fmri] Missing 390 functional scans for session 1.
[fetch_spm_multimodal_fmri] Data absent, downloading...
[fetch_spm_multimodal_fmri] Downloading data from
https://www.fil.ion.ucl.ac.uk/spm/download/data/mmfaces/multimodal_fmri.zip ...
[fetch_spm_multimodal_fmri] Downloaded 581632 of 134263085 bytes (0.4%%, 00 HR
03 MIN 53 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 4661248 of 134263085 bytes (3.5%%, 00 HR
00 MIN 57 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 11116544 of 134263085 bytes (8.3%%, 00 HR
00 MIN 34 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 18153472 of 134263085 bytes (13.5%%, 00
HR 00 MIN 26 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 25255936 of 134263085 bytes (18.8%%, 00
HR 00 MIN 22 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 30842880 of 134263085 bytes (23.0%%, 00
HR 00 MIN 21 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 36782080 of 134263085 bytes (27.4%%, 00
HR 00 MIN 19 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 42819584 of 134263085 bytes (31.9%%, 00
HR 00 MIN 17 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 47382528 of 134263085 bytes (35.3%%, 00
HR 00 MIN 17 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 52256768 of 134263085 bytes (38.9%%, 00
HR 00 MIN 16 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 57352192 of 134263085 bytes (42.7%%, 00
HR 00 MIN 15 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 62570496 of 134263085 bytes (46.6%%, 00
HR 00 MIN 14 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 67854336 of 134263085 bytes (50.5%%, 00
HR 00 MIN 13 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 73154560 of 134263085 bytes (54.5%%, 00
HR 00 MIN 12 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 78462976 of 134263085 bytes (58.4%%, 00
HR 00 MIN 11 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 83763200 of 134263085 bytes (62.4%%, 00
HR 00 MIN 10 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 88727552 of 134263085 bytes (66.1%%, 00
HR 00 MIN 09 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 92831744 of 134263085 bytes (69.1%%, 00
HR 00 MIN 08 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 97386496 of 134263085 bytes (72.5%%, 00
HR 00 MIN 07 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 102277120 of 134263085 bytes (76.2%%, 00
HR 00 MIN 06 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 107388928 of 134263085 bytes (80.0%%, 00
HR 00 MIN 05 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 112631808 of 134263085 bytes (83.9%%, 00
HR 00 MIN 04 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 117948416 of 134263085 bytes (87.8%%, 00
HR 00 MIN 03 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 123289600 of 134263085 bytes (91.8%%, 00
HR 00 MIN 02 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 128638976 of 134263085 bytes (95.8%%, 00
HR 00 MIN 01 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 133980160 of 134263085 bytes (99.8%%, 00
HR 00 MIN 00 SEC remaining)
[fetch_spm_multimodal_fmri] ...done. (28 seconds, 0 min)
[fetch_spm_multimodal_fmri] Extracting data from
/home/runner/nilearn_data/spm_multimodal_fmri/sub001/multimodal_fmri.zip...
[fetch_spm_multimodal_fmri] .. done.
[fetch_spm_multimodal_fmri] Downloading data from
https://www.fil.ion.ucl.ac.uk/spm/download/data/mmfaces/multimodal_smri.zip ...
[fetch_spm_multimodal_fmri] Downloaded 557056 of 6852766 bytes (8.1%%, 00 HR 00
MIN 12 SEC remaining)
[fetch_spm_multimodal_fmri] Downloaded 4145152 of 6852766 bytes (60.5%%, 00 HR
00 MIN 01 SEC remaining)
[fetch_spm_multimodal_fmri] ...done. (3 seconds, 0 min)
[fetch_spm_multimodal_fmri] Extracting data from
/home/runner/nilearn_data/spm_multimodal_fmri/sub001/multimodal_smri.zip...
[fetch_spm_multimodal_fmri] .. done.
Let’s inspect one of the event files before using them.
import pandas as pd
events = [subject_data.events1, subject_data.events2]
events_dataframe = pd.read_csv(events[0], sep="\t")
events_dataframe["trial_type"].value_counts()
trial_type
scrambled 86
faces 64
Name: count, dtype: int64
We can confirm there are only 2 conditions in the dataset.
from nilearn.plotting import plot_event, show
plot_event(events)
show()
Resample the images:
this is achieved by the concat_imgs function of Nilearn.
import warnings
from nilearn.image import concat_imgs, mean_img, resample_img
# Avoid getting too many warnings due to resampling
with warnings.catch_warnings():
warnings.simplefilter("ignore")
fmri_img = [
concat_imgs(subject_data.func1, auto_resample=True),
concat_imgs(subject_data.func2, auto_resample=True),
]
affine, shape = fmri_img[0].affine, fmri_img[0].shape
print("Resampling the second image (this takes time)...")
fmri_img[1] = resample_img(fmri_img[1], affine, shape[:3])
Resampling the second image (this takes time)...
Let’s create mean image for display purposes.
Fit the model¶
Fit the GLM for the 2 runs by specifying a FirstLevelModel and then fitting it.
# Sample at the beginning of each acquisition.
slice_time_ref = 0.0
# We use a discrete cosine transform to model signal drifts.
drift_model = "cosine"
# The cutoff for the drift model is 0.01 Hz.
high_pass = 0.01
# The hemodynamic response function
hrf_model = "spm + derivative"
from nilearn.glm.first_level import FirstLevelModel
print("Fitting a GLM")
fmri_glm = FirstLevelModel(
smoothing_fwhm=None,
t_r=subject_data.t_r,
hrf_model=hrf_model,
drift_model=drift_model,
high_pass=high_pass,
verbose=1,
)
fmri_glm = fmri_glm.fit(fmri_img, events=events)
Fitting a GLM
\[FirstLevelModel.fit] Computing run 1 out of 2 runs (go take a coffee, a big
one).
\[FirstLevelModel.fit] Performing mask computation.
\[FirstLevelModel.fit] Masking took 0 seconds.
\[FirstLevelModel.fit] Performing GLM computation.
\[FirstLevelModel.fit] GLM took 1 seconds.
\[FirstLevelModel.fit] Computing run 2 out of 2 runs (00 HR 00 MIN 02 SEC
remaining).
\[FirstLevelModel.fit] Performing mask computation.
\[FirstLevelModel.fit] Masking took 0 seconds.
\[FirstLevelModel.fit] Performing GLM computation.
\[FirstLevelModel.fit] GLM took 0 seconds.
\[FirstLevelModel.fit] Computation of 2 runs done in 00 HR 00 MIN 04 SEC.
View the results¶
Now we can compute contrast-related statistical maps (in z-scale), and plot them.
from nilearn.plotting import plot_stat_map
print("Computing contrasts")
Computing contrasts
We actually want more interesting contrasts. The simplest contrast just makes the difference between the two main conditions. We define the two opposite versions to run one-tailed t-tests.
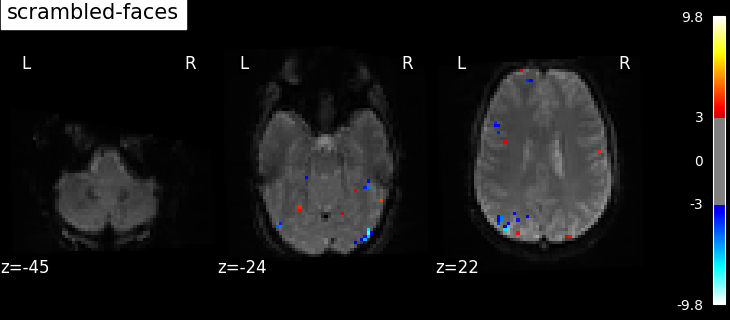
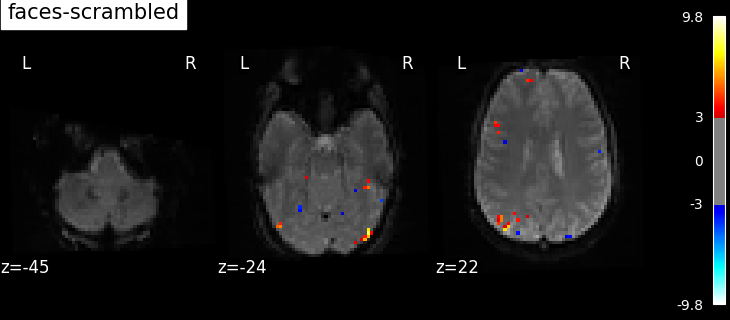
contrasts = ["faces - scrambled", "scrambled - faces"]
Let’s store common parameters for all plots.
We plot the contrasts values overlaid on the mean fMRI image and we will use the z-score values as transparency, with any voxel with | Z-score | > 3 being fully opaque and any voxel with 0 < | Z-score | < 1.96 being partly transparent.
plot_param = {
"vmin": 0,
"display_mode": "z",
"cut_coords": 3,
"black_bg": True,
"bg_img": mean_image,
"cmap": "inferno",
"transparency_range": [0, 3],
}
# Iterate on contrasts to compute and plot them.
for contrast_id in contrasts:
print(f"\tcontrast id: {contrast_id}")
results = fmri_glm.compute_contrast(contrast_id, output_type="all")
plot_stat_map(
results["stat"],
title=contrast_id,
transparency=results["z_score"],
**plot_param,
)
contrast id: faces - scrambled
/home/runner/work/nilearn/nilearn/examples/04_glm_first_level/plot_spm_multimodal_faces.py:134: RuntimeWarning:
The same contrast will be used for all 2 runs. If the design matrices are not the same for all runs, (for example with different column names or column order across runs) you should pass contrast as an expression using the name of the conditions as they appear in the design matrices.
contrast id: scrambled - faces
/home/runner/work/nilearn/nilearn/examples/04_glm_first_level/plot_spm_multimodal_faces.py:134: RuntimeWarning:
The same contrast will be used for all 2 runs. If the design matrices are not the same for all runs, (for example with different column names or column order across runs) you should pass contrast as an expression using the name of the conditions as they appear in the design matrices.
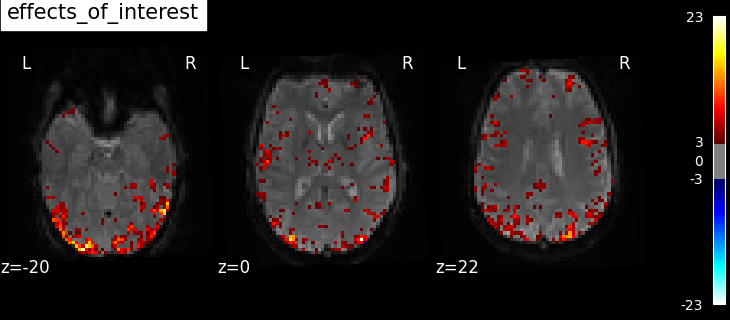
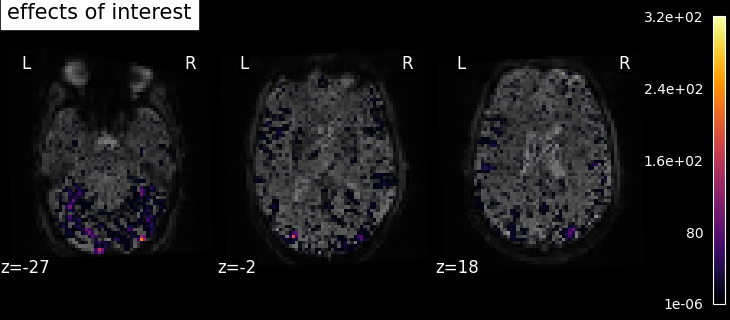
We also define the effects of interest contrast, a 2-dimensional contrasts spanning the two conditions.
import numpy as np
contrasts = np.eye(2)
results = fmri_glm.compute_contrast(contrasts, output_type="all")
plot_stat_map(
results["stat"],
title="effects of interest",
transparency=results["z_score"],
**plot_param,
)
show()
/home/runner/work/nilearn/nilearn/examples/04_glm_first_level/plot_spm_multimodal_faces.py:151: RuntimeWarning:
The same contrast will be used for all 2 runs. If the design matrices are not the same for all runs, (for example with different column names or column order across runs) you should pass contrast as an expression using the name of the conditions as they appear in the design matrices.
/home/runner/work/nilearn/nilearn/examples/04_glm_first_level/plot_spm_multimodal_faces.py:151: UserWarning:
F contrasts should have 20 columns, but it has only 2. The rest of the contrast was padded with zeros.
/home/runner/work/nilearn/nilearn/examples/04_glm_first_level/plot_spm_multimodal_faces.py:151: UserWarning:
Running approximate fixed effects on F statistics.
Based on the resulting maps we observe that the analysis results in wide activity for the ‘effects of interest’ contrast, showing the implications of large portions of the visual cortex in the conditions. By contrast, the differential effect between “faces” and “scrambled” involves sparser, more anterior and lateral regions. It also displays some responses in the frontal lobe.
Total running time of the script: (1 minutes 30.435 seconds)
Estimated memory usage: 963 MB