Note
This page is a reference documentation. It only explains the class signature, and not how to use it. Please refer to the user guide for the big picture.
nilearn.regions.RegionExtractor¶
- class nilearn.regions.RegionExtractor(maps_img=None, mask_img=None, min_region_size=1350, threshold=1.0, thresholding_strategy='ratio_n_voxels', two_sided=False, extractor='local_regions', smoothing_fwhm=6, standardize=False, standardize_confounds=True, high_variance_confounds=False, detrend=False, low_pass=None, high_pass=None, t_r=None, dtype=None, resampling_target='data', keep_masked_maps=False, memory=None, memory_level=0, verbose=0, reports=True, cmap='CMRmap_r', allow_overlap=True, clean_args=None)[source]¶
Class for brain region extraction.
Region Extraction is a post processing technique which is implemented to automatically segment each brain atlas maps into different set of separated brain activated region. Particularly, to show that each decomposed brain maps can be used to focus on a target specific Regions of Interest analysis.
See Abraham et al.[1].
Added in Nilearn 0.2.
- Parameters:
- maps_img4D Niimg-like object or None, default=None
Image containing a set of whole brain atlas maps or statistically decomposed brain maps.
- mask_imgNiimg-like object or None, default=None
Mask to be applied to input data, passed to NiftiMapsMasker. If None, no masking is applied.
- min_region_size
float, default=1350 Minimum volume in mm3 for a region to be kept. For example, if the voxel size is 3x3x3 mm then the volume of the voxel is 27mm^3. The default of 1350mm^3 means we take minimum size of 1350 / 27 = 50 voxels.
- thresholdnumber, default=1.0
A value used either in ratio_n_voxels or img_value or percentile thresholding_strategy based upon the choice of selection.
- thresholding_strategy
str{‘ratio_n_voxels’, ‘img_value’, ‘percentile’}, default=’ratio_n_voxels’ If default ‘ratio_n_voxels’, we apply thresholding that will keep the more intense nonzero brain voxels (denoted as n_voxels) across all maps (n_voxels being the number of voxels in the brain volume). A float value given in threshold parameter indicates the ratio of voxels to keep meaning (if float=2. then maps will together have 2. x n_voxels non-zero voxels). If set to ‘percentile’, images are thresholded based on the score obtained with the given percentile on the data and the voxel intensities which are survived above this obtained score will be kept. If set to ‘img_value’, we apply thresholding based on the non-zero voxel intensities across all maps. A value given in threshold parameter indicates that we keep only those voxels which have intensities more than this value.
- two_sided
bool, default=False Whether the thresholding should yield both positive and negative part of the maps.
Added in Nilearn 0.11.1.
- extractor{“local_regions”, “connected_components”}, default=”local_regions”
This option can take two values:
"connected_components": each component/region in the image is extracted automatically by labeling each region based upon the presence of unique features in their respective regions."local_regions": each component/region is extracted based on their maximum peak value to define a seed marker and then using random walker segmentation algorithm on these markers for region separation.
- smoothing_fwhm
floatorintor None, optional. If smoothing_fwhm is not None, it gives the full-width at half maximum in millimeters of the spatial smoothing to apply to the signal. Use this parameter to smooth an image to extract most sparser regions.
Note
This parameter is passed to
nilearn.regions.connected_regions. It will be used only ifextractor='local_regions'.Note
Please set this parameter according to maps resolution, otherwise extraction will fail.
default=6mm.
- standardizeany of: ‘zscore_sample’, ‘zscore’, ‘psc’, True, False or None; default=False
Strategy to standardize the signal:
'zscore_sample': The signal is z-scored. Timeseries are shifted to zero mean and scaled to unit variance. Uses sample std.'psc': Timeseries are shifted to zero mean value and scaled to percent signal change (as compared to original mean signal).True: The signal is z-scored (same as option zscore). Timeseries are shifted to zero mean and scaled to unit variance.Deprecated since Nilearn 0.13.0: In nilearn version 0.15.0,
Truewill be replaced by'zscore_sample'.False: Do not standardize the data.Deprecated since Nilearn 0.13.0: In nilearn version 0.15.0,
Falsewill be replaced byNone.
Deprecated since Nilearn 0.13.0: The default will be changed to
Nonein version 0.15.0.Note
Recommended to set to True if signals are not already standardized. Passed to
NiftiMapsMasker.- standardize_confounds
bool, default=True If set to True, the confounds are z-scored: their mean is put to 0 and their variance to 1 in the time dimension.
- high_variance_confounds
bool, default=False If True, high variance confounds are computed on provided image with
nilearn.image.high_variance_confoundsand default parameters and regressed out.- detrend
bool, optional Whether to detrend signals or not.
Note
Passed to
nilearn.signal.clean.default=False.
- low_pass
floatorintor None, default=None Low cutoff frequency in Hertz. If specified, signals above this frequency will be filtered out. If None, no low-pass filtering will be performed.
Note
Passed to
nilearn.signal.clean.- high_pass
floatorintor None, default=None High cutoff frequency in Hertz. If specified, signals below this frequency will be filtered out.
Note
Passed to
nilearn.signal.clean.- t_r
floatorintor None, default=None Repetition time, in seconds (sampling period). Set to None if not provided.
Note
Passed to
nilearn.signal.clean.- dtypedtype like, “auto” or None, default=None
Data type toward which the data should be converted. If “auto”, the data will be converted to int32 if dtype is discrete and float32 if it is continuous. If None, data will not be converted to a new data type.
- resampling_target{“data”, “mask”, “maps”, None}, default=”data”
Defines which image gives the final shape/size.
"data"means that the atlas is resampled to the shape of the data if needed"mask"means that themaps_imgand images provided tofit()are resampled to the shape and affine ofmask_img"maps"means themask_imgand images provided tofit()are resampled to the shape and affine ofmaps_imgNonemeans no resampling: if shapes and affines do not match, aValueErroris raised.
- keep_masked_maps
bool, optional If True, masked atlas with invalid maps (maps that contain only zeros after applying the mask) will be retained in the output, resulting in corresponding time series containing zeros only. If False, the invalid maps will be removed from the trimmed atlas, resulting in no empty time series in the output.
Deprecated since Nilearn 0.10.2.
Changed in Nilearn 0.13.0: The
keep_masked_mapsparameter will be removed in 0.15.- memoryNone, instance of
joblib.Memory,str, orpathlib.Path, default=None Used to cache the masking process. By default, no caching is done. If a
stris given, it is the path to the caching directory.- memory_level
int, default=0 Rough estimator of the amount of memory used by caching. Higher value means more memory for caching. Zero means no caching.
- verbose
boolorint, default=0 Verbosity level (
0orFalsemeans no message).- reports
bool, default=True If set to True, data is saved in order to produce a report.
- cmap
matplotlib.colors.Colormap, orstr, optional The colormap to use. Either a string which is a name of a matplotlib colormap, or a matplotlib colormap object. default=”CMRmap_r” Only relevant for the report figures.
- allow_overlapTrue
If False, an error is raised if the maps overlaps (ie at least two maps have a non-zero value for the same voxel).
- clean_args
dictor None, default=None Keyword arguments to be passed to
cleancalled within the masker. Withinclean, kwargs prefixed with'butterworth__'will be passed to the Butterworth filter.Added in Nilearn 0.12.1.
- Attributes:
- clean_args_
dict Keyword arguments to be passed to
cleancalled within the masker. Withinclean, kwargs prefixed with'butterworth__'will be passed to the Butterworth filter.- index_
numpy.ndarray Array of list of indices where each index value is assigned to each separate region of its corresponding family of brain maps.
- maps_img_
nibabel.nifti1.Nifti1Image The maps mask of the data.
- mask_img_A 3D binary
nibabel.nifti1.Nifti1Imageor None. The mask of the data. If no
mask_imgwas passed at masker construction, thenmask_img_isNone, otherwise is the resulting binarized version ofmask_imgwhere each voxel isTrueif all values across samples (for example across timepoints) is finite value different from 0.- memory_joblib memory cache
- n_elements_
int The number of overlapping maps in the mask. This is equivalent to the number of volumes in the mask image.
Added in Nilearn 0.9.2.
- regions_img_
nibabel.nifti1.Nifti1Image List of separated regions with each region lying on an original volume concatenated into a 4D image.
- clean_args_
See also
nilearn.regions.connected_label_regionsA function can be readily used for extraction of regions on labels based atlas images.
References
- __init__(maps_img=None, mask_img=None, min_region_size=1350, threshold=1.0, thresholding_strategy='ratio_n_voxels', two_sided=False, extractor='local_regions', smoothing_fwhm=6, standardize=False, standardize_confounds=True, high_variance_confounds=False, detrend=False, low_pass=None, high_pass=None, t_r=None, dtype=None, resampling_target='data', keep_masked_maps=False, memory=None, memory_level=0, verbose=0, reports=True, cmap='CMRmap_r', allow_overlap=True, clean_args=None)[source]¶
- fit(imgs=None, y=None)[source]¶
Prepare signal extraction from regions.
- Parameters:
- imgs
listof Niimg-like objects or None, default=None See Input and output: neuroimaging data representation. Image data passed to the reporter.
- yNone
This parameter is unused. It is solely included for scikit-learn compatibility.
- imgs
- fit_transform(imgs, y=None, confounds=None, sample_mask=None)[source]¶
Prepare and perform signal extraction.
- Parameters:
- imgs3D/4D Niimg-like object
See Input and output: neuroimaging data representation. Images to process. If a 3D niimg is provided, a 1D array is returned.
- yNone
This parameter is unused. It is solely included for scikit-learn compatibility.
- confounds
numpy.ndarray,str,pathlib.Path,pandas.DataFrameorlistof confounds timeseries, default=None This parameter is passed to
nilearn.signal.clean. Please see the related documentation for details. shape: (number of scans, number of confounds)- sample_maskAny type compatible with numpy-array indexing, default=None
shape = (total number of scans - number of scans removed)for explicit index (for example,sample_mask=np.asarray([1, 2, 4])), orshape = (number of scans)for binary mask (for example,sample_mask=np.asarray([False, True, True, False, True])). Masks the images along the last dimension to perform scrubbing: for example to remove volumes with high motion and/or non-steady-state volumes. This parameter is passed tonilearn.signal.clean.Added in Nilearn 0.8.0.
- Returns:
- signals
numpy.ndarray,pandas.DataFrameor polars.DataFrame Signal for each element.
Changed in Nilearn 0.13.0: Added
set_outputsupport.The type of the output is determined by
set_output(): see the scikit-learn documentation.Output shape for :
For Numpy outputs:
3D images: (number of elements,)
4D images: (number of scans, number of elements) array
For DataFrame outputs:
3D or 4D images: (number of scans, number of elements) array
- signals
- generate_report(engine='matplotlib', title=None, **kwargs)[source]¶
Generate an HTML report for this masker.
The
HTMLReportcan be opened in a browser, displayed in a notebook, or saved to disk as a standalone HTML file.Note
This functionality requires to have
Matplotlibinstalled.- Parameters:
- engine
str, default=”matplotlib” Choice of engine to display report figures.
All maskers support
"matplotlib"as engine.Other options are :
"brainsprite":NiftiMasker, MultiNiftiMasker, NiftiLabelsMasker, MultiNiftiLabelsMasker:
"plotly":SurfaceMasker, MultiSurfaceMasker, SurfaceLabelsMasker, MultiSurfaceLabelsMasker, SurfaceMapsMasker, MultiSurfaceMapsMasker:
- title
stror None, default=None title for the report. If None, title will be the class name.
- kwargs
dict[str, Any] Dictionary of key-word arguments necessary for report generation.
Expected keys depending on masker type are:
For NiftiMapsMasker, MultiNiftiMapsMasker, SurfaceMapsMasker, MultiSurfaceMapsMasker :
- displayed_maps
int,ndarrayorlistofint, or “all”, default=10 Indicates which maps will be displayed in the HTML report.
If
"all": All maps will be displayed in the report.
masker.generate_report("all")
masker.generate_report([6, 3, 12])
- If an
int: This will only display the first n maps, n being the value of the parameter. By default, the report will only contain the first 10 maps. Example to display the first 16 maps:
- If an
masker.generate_report(16)
For NiftiSpheresMasker :
- displayed_spheres
int,ndarrayorlistofint, or “all”, default=10 Indicates which spheres will be displayed in the HTML report.
If
"all": All spheres will be displayed in the report.
masker.generate_report("all")
masker.generate_report([6, 3, 12])
- If an
int: This will only display the first n spheres, n being the value of the parameter. By default, the report will only contain the first 10 spheres. Example to display the first 16 spheres:
- If an
masker.generate_report(16)
- engine
- Returns:
- reportnilearn.reporting.HTMLReport
HTML report for the masker.
- get_feature_names_out(input_features=None)¶
Get output feature names for transformation.
The feature names out will prefixed by the lowercased class name. For example, if the transformer outputs 3 features, then the feature names out are: [“class_name0”, “class_name1”, “class_name2”].
- Parameters:
- input_featuresarray-like of str or None, default=None
Only used to validate feature names with the names seen in fit.
- Returns:
- feature_names_outndarray of str objects
Transformed feature names.
- get_metadata_routing()¶
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)¶
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- inverse_transform(region_signals)[source]¶
Compute voxel signals from region signals.
Any mask given at initialization is taken into account.
- Parameters:
- signals1D/2D
numpy.ndarray Extracted signal. If a 1D array is provided, then the shape should be (number of elements,). If a 2D array is provided, then the shape should be (number of scans, number of elements).
- signals1D/2D
- Returns:
- img
nibabel.nifti1.Nifti1Image Transformed image in brain space. Output shape for :
1D array : 3D
nibabel.nifti1.Nifti1Imagewill be returned.2D array : 4D
nibabel.nifti1.Nifti1Imagewill be returned.
- img
- set_fit_request(*, imgs='$UNCHANGED$')¶
Request metadata passed to the
fitmethod.Note that this method is only relevant if
enable_metadata_routing=True(seesklearn.set_config). Please see User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed tofitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it tofit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
Note
This method is only relevant if this estimator is used as a sub-estimator of a meta-estimator, e.g. used inside a
Pipeline. Otherwise it has no effect.- Parameters:
- imgsstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
imgsparameter infit.
- Returns:
- selfobject
The updated object.
- set_inverse_transform_request(*, region_signals='$UNCHANGED$')¶
Request metadata passed to the
inverse_transformmethod.Note that this method is only relevant if
enable_metadata_routing=True(seesklearn.set_config). Please see User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed toinverse_transformif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it toinverse_transform.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
Note
This method is only relevant if this estimator is used as a sub-estimator of a meta-estimator, e.g. used inside a
Pipeline. Otherwise it has no effect.- Parameters:
- region_signalsstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
region_signalsparameter ininverse_transform.
- Returns:
- selfobject
The updated object.
- set_output(*, transform=None)¶
Set output container.
See Introducing the set_output API for an example on how to use the API.
- Parameters:
- transform{“default”, “pandas”, “polars”}, default=None
Configure output of transform and fit_transform.
“default”: Default output format of a transformer
“pandas”: DataFrame output
“polars”: Polars output
None: Transform configuration is unchanged
Added in version 1.4: “polars” option was added.
- Returns:
- selfestimator instance
Estimator instance.
- set_params(**params)¶
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- set_transform_request(*, confounds='$UNCHANGED$', imgs='$UNCHANGED$', sample_mask='$UNCHANGED$')¶
Request metadata passed to the
transformmethod.Note that this method is only relevant if
enable_metadata_routing=True(seesklearn.set_config). Please see User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed totransformif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it totransform.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
Note
This method is only relevant if this estimator is used as a sub-estimator of a meta-estimator, e.g. used inside a
Pipeline. Otherwise it has no effect.- Parameters:
- confoundsstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
confoundsparameter intransform.- imgsstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
imgsparameter intransform.- sample_maskstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_maskparameter intransform.
- Returns:
- selfobject
The updated object.
- transform(imgs, confounds=None, sample_mask=None)[source]¶
Apply mask, spatial and temporal preprocessing.
- Parameters:
- imgs3D/4D Niimg-like object
See Input and output: neuroimaging data representation. Images to process. If a 3D niimg is provided, a 1D array is returned.
- confounds
numpy.ndarray,str,pathlib.Path,pandas.DataFrameorlistof confounds timeseries, default=None This parameter is passed to
nilearn.signal.clean. Please see the related documentation for details. shape: (number of scans, number of confounds)- sample_maskAny type compatible with numpy-array indexing, default=None
shape = (total number of scans - number of scans removed)for explicit index (for example,sample_mask=np.asarray([1, 2, 4])), orshape = (number of scans)for binary mask (for example,sample_mask=np.asarray([False, True, True, False, True])). Masks the images along the last dimension to perform scrubbing: for example to remove volumes with high motion and/or non-steady-state volumes. This parameter is passed tonilearn.signal.clean.Added in Nilearn 0.8.0.
- Returns:
- signals
numpy.ndarray,pandas.DataFrameor polars.DataFrame Signal for each element.
Changed in Nilearn 0.13.0: Added
set_outputsupport.The type of the output is determined by
set_output(): see the scikit-learn documentation.Output shape for :
For Numpy outputs:
3D images: (number of elements,)
4D images: (number of scans, number of elements) array
For DataFrame outputs:
3D or 4D images: (number of scans, number of elements) array
- signals
- transform_single_imgs(imgs, confounds=None, sample_mask=None)[source]¶
Extract signals from a single 4D niimg.
- Parameters:
- imgs3D/4D Niimg-like object
See Input and output: neuroimaging data representation. Images to process.
- confoundsCSV file or array-like, default=None
This parameter is passed to
nilearn.signal.clean. Please see the related documentation for details. shape: (number of scans, number of confounds)- sample_maskAny type compatible with numpy-array indexing, default=None
shape = (total number of scans - number of scans removed)for explicit index (for example,sample_mask=np.asarray([1, 2, 4])), orshape = (number of scans)for binary mask (for example,sample_mask=np.asarray([False, True, True, False, True])). Masks the images along the last dimension to perform scrubbing: for example to remove volumes with high motion and/or non-steady-state volumes. This parameter is passed tonilearn.signal.clean.Added in Nilearn 0.8.0.
- Returns:
- signals
numpy.ndarray,pandas.DataFrameor polars.DataFrame Signal for each element.
Changed in Nilearn 0.13.0: Added
set_outputsupport.The type of the output is determined by
set_output(): see the scikit-learn documentation.Output shape for :
For Numpy outputs:
3D images: (number of elements,)
4D images: (number of scans, number of elements) array
For DataFrame outputs:
3D or 4D images: (number of scans, number of elements) array
- signals
Examples using nilearn.regions.RegionExtractor¶
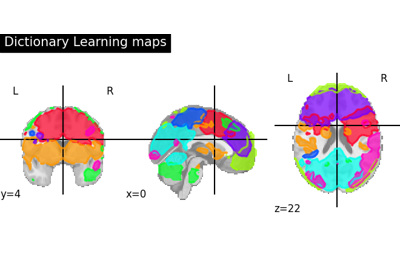
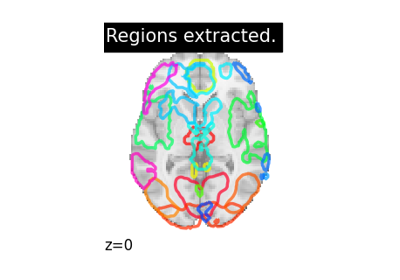
Regions extraction using dictionary learning and functional connectomes
Regions Extraction of Default Mode Networks using Smith Atlas