Note
This page is a reference documentation. It only explains the class signature, and not how to use it. Please refer to the user guide for the big picture.
nilearn.maskers.BaseMasker¶
- class nilearn.maskers.BaseMasker[source]¶
Base class for NiftiMaskers.
- fit(imgs=None, y=None)[source]¶
Compute the mask corresponding to the data.
- Parameters:
- imgs
listof Niimg-like objects or None, default=None See Input and output: neuroimaging data representation. Data on which the mask must be calculated. If this is a list, the affine is considered the same for all.
- yNone
This parameter is unused. It is solely included for scikit-learn compatibility.
- imgs
- fit_transform(imgs, y=None, confounds=None, sample_mask=None, **fit_params)[source]¶
Fit to data, then transform it.
- Parameters:
- imgsNiimg-like object
- yNone
This parameter is unused. It is solely included for scikit-learn compatibility.
- confounds
numpy.ndarray,str,pathlib.Path,pandas.DataFrameorlistof confounds timeseries, default=None This parameter is passed to
nilearn.signal.clean. Please see the related documentation for details. shape: (number of scans, number of confounds)- sample_maskAny type compatible with numpy-array indexing, default=None
shape = (total number of scans - number of scans removed)for explicit index (for example,sample_mask=np.asarray([1, 2, 4])), orshape = (number of scans)for binary mask (for example,sample_mask=np.asarray([False, True, True, False, True])). Masks the images along the last dimension to perform scrubbing: for example to remove volumes with high motion and/or non-steady-state volumes. This parameter is passed tonilearn.signal.clean.Added in Nilearn 0.8.0.
- Returns:
- signals
numpy.ndarray,pandas.DataFrameor polars.DataFrame Signal for each element.
Changed in Nilearn 0.13.0: Added
set_outputsupport.The type of the output is determined by
set_output(): see the scikit-learn documentation.Output shape for :
For Numpy outputs:
3D images: (number of elements,)
4D images: (number of scans, number of elements) array
For DataFrame outputs:
3D or 4D images: (number of scans, number of elements) array
- signals
- generate_report(engine='matplotlib', title=None, **kwargs)[source]¶
Generate an HTML report for this masker.
The
HTMLReportcan be opened in a browser, displayed in a notebook, or saved to disk as a standalone HTML file.Note
This functionality requires to have
Matplotlibinstalled.- Parameters:
- engine
str, default=”matplotlib” Choice of engine to display report figures.
All maskers support
"matplotlib"as engine.Other options are :
"brainsprite":NiftiMasker, MultiNiftiMasker, NiftiLabelsMasker, MultiNiftiLabelsMasker:
"plotly":SurfaceMasker, MultiSurfaceMasker, SurfaceLabelsMasker, MultiSurfaceLabelsMasker, SurfaceMapsMasker, MultiSurfaceMapsMasker:
- title
stror None, default=None title for the report. If None, title will be the class name.
- kwargs
dict[str, Any] Dictionary of key-word arguments necessary for report generation.
Expected keys depending on masker type are:
For NiftiMapsMasker, MultiNiftiMapsMasker, SurfaceMapsMasker, MultiSurfaceMapsMasker :
- displayed_maps
int,ndarrayorlistofint, or “all”, default=10 Indicates which maps will be displayed in the HTML report.
If
"all": All maps will be displayed in the report.
masker.generate_report("all")
masker.generate_report([6, 3, 12])
- If an
int: This will only display the first n maps, n being the value of the parameter. By default, the report will only contain the first 10 maps. Example to display the first 16 maps:
- If an
masker.generate_report(16)
For NiftiSpheresMasker :
- displayed_spheres
int,ndarrayorlistofint, or “all”, default=10 Indicates which spheres will be displayed in the HTML report.
If
"all": All spheres will be displayed in the report.
masker.generate_report("all")
masker.generate_report([6, 3, 12])
- If an
int: This will only display the first n spheres, n being the value of the parameter. By default, the report will only contain the first 10 spheres. Example to display the first 16 spheres:
- If an
masker.generate_report(16)
- engine
- Returns:
- reportnilearn.reporting.HTMLReport
HTML report for the masker.
- get_metadata_routing()¶
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)¶
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- inverse_transform(X)[source]¶
Transform the data matrix back to an image in brain space.
This step only performs spatial unmasking, without inverting any additional processing performed by
transform, such as temporal filtering or smoothing.- Parameters:
- signals1D/2D
numpy.ndarray Extracted signal. If a 1D array is provided, then the shape should be (number of elements,). If a 2D array is provided, then the shape should be (number of scans, number of elements).
- signals1D/2D
- Returns:
- img
nibabel.nifti1.Nifti1Image Transformed image in brain space. Output shape for :
1D array : 3D
nibabel.nifti1.Nifti1Imagewill be returned.2D array : 4D
nibabel.nifti1.Nifti1Imagewill be returned.
- img
- set_fit_request(*, imgs='$UNCHANGED$')¶
Request metadata passed to the
fitmethod.Note that this method is only relevant if
enable_metadata_routing=True(seesklearn.set_config). Please see User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed tofitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it tofit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
Note
This method is only relevant if this estimator is used as a sub-estimator of a meta-estimator, e.g. used inside a
Pipeline. Otherwise it has no effect.- Parameters:
- imgsstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
imgsparameter infit.
- Returns:
- selfobject
The updated object.
- set_output(*, transform=None)¶
Set output container.
See Introducing the set_output API for an example on how to use the API.
- Parameters:
- transform{“default”, “pandas”, “polars”}, default=None
Configure output of transform and fit_transform.
“default”: Default output format of a transformer
“pandas”: DataFrame output
“polars”: Polars output
None: Transform configuration is unchanged
Added in version 1.4: “polars” option was added.
- Returns:
- selfestimator instance
Estimator instance.
- set_params(**params)¶
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- set_transform_request(*, confounds='$UNCHANGED$', imgs='$UNCHANGED$', sample_mask='$UNCHANGED$')¶
Request metadata passed to the
transformmethod.Note that this method is only relevant if
enable_metadata_routing=True(seesklearn.set_config). Please see User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed totransformif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it totransform.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
Note
This method is only relevant if this estimator is used as a sub-estimator of a meta-estimator, e.g. used inside a
Pipeline. Otherwise it has no effect.- Parameters:
- confoundsstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
confoundsparameter intransform.- imgsstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
imgsparameter intransform.- sample_maskstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_maskparameter intransform.
- Returns:
- selfobject
The updated object.
- transform(imgs, confounds=None, sample_mask=None)[source]¶
Apply mask, spatial and temporal preprocessing.
- Parameters:
- imgs3D/4D Niimg-like object
See Input and output: neuroimaging data representation. Images to process. If a 3D niimg is provided, a 1D array is returned.
- confounds
numpy.ndarray,str,pathlib.Path,pandas.DataFrameorlistof confounds timeseries, default=None This parameter is passed to
nilearn.signal.clean. Please see the related documentation for details. shape: (number of scans, number of confounds)- sample_maskAny type compatible with numpy-array indexing, default=None
shape = (total number of scans - number of scans removed)for explicit index (for example,sample_mask=np.asarray([1, 2, 4])), orshape = (number of scans)for binary mask (for example,sample_mask=np.asarray([False, True, True, False, True])). Masks the images along the last dimension to perform scrubbing: for example to remove volumes with high motion and/or non-steady-state volumes. This parameter is passed tonilearn.signal.clean.Added in Nilearn 0.8.0.
- Returns:
- signals
numpy.ndarray,pandas.DataFrameor polars.DataFrame Signal for each element.
Changed in Nilearn 0.13.0: Added
set_outputsupport.The type of the output is determined by
set_output(): see the scikit-learn documentation.Output shape for :
For Numpy outputs:
3D images: (number of elements,)
4D images: (number of scans, number of elements) array
For DataFrame outputs:
3D or 4D images: (number of scans, number of elements) array
- signals
- abstract transform_single_imgs(imgs, confounds=None, sample_mask=None, copy=True)[source]¶
Extract signals from a single niimg.
- Parameters:
- imgs3D/4D Niimg-like object
See Input and output: neuroimaging data representation. Images to process.
- confounds
numpy.ndarray,str,pathlib.Path,pandas.DataFrameorlistof confounds timeseries, default=None This parameter is passed to
nilearn.signal.clean. Please see the related documentation for details. shape: (number of scans, number of confounds)- sample_maskAny type compatible with numpy-array indexing, default=None
shape = (total number of scans - number of scans removed)for explicit index (for example,sample_mask=np.asarray([1, 2, 4])), orshape = (number of scans)for binary mask (for example,sample_mask=np.asarray([False, True, True, False, True])). Masks the images along the last dimension to perform scrubbing: for example to remove volumes with high motion and/or non-steady-state volumes. This parameter is passed tonilearn.signal.clean.Added in Nilearn 0.8.0.
- copy
bool, default=True Indicates whether a copy is returned or not.
- Returns:
- signals
numpy.ndarray,pandas.DataFrameor polars.DataFrame Signal for each element.
Changed in Nilearn 0.13.0: Added
set_outputsupport.The type of the output is determined by
set_output(): see the scikit-learn documentation.Output shape for :
For Numpy outputs:
3D images: (number of elements,)
4D images: (number of scans, number of elements) array
For DataFrame outputs:
3D or 4D images: (number of scans, number of elements) array
- signals
Examples using nilearn.maskers.BaseMasker¶

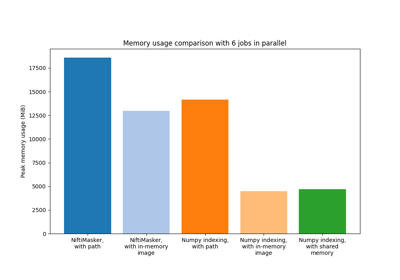
Encoding models for visual stimuli from Miyawaki et al. 2008
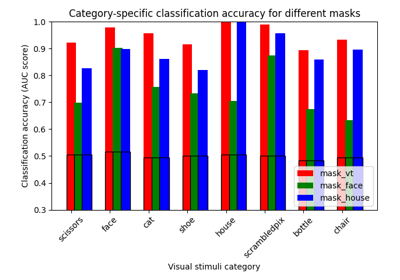
ROI-based decoding analysis in Haxby et al. dataset
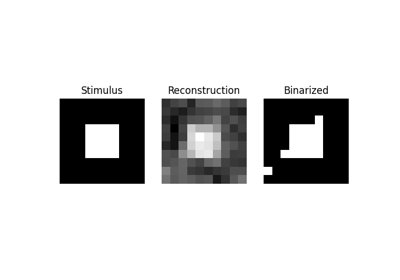

Reconstruction of visual stimuli from Miyawaki et al. 2008
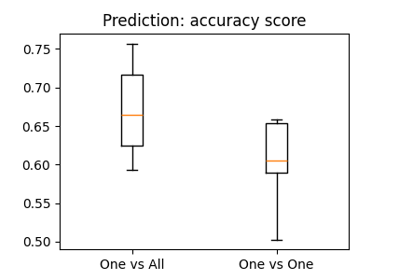
The haxby dataset: different multi-class strategies
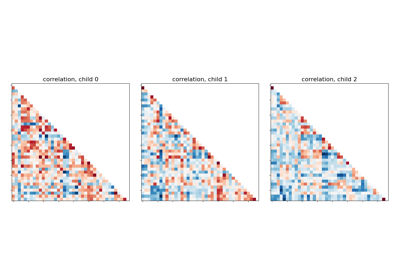
Classification of age groups using functional connectivity
Comparing connectomes on different reference atlases
Computing a connectome with sparse inverse covariance
Extracting signals of a probabilistic atlas of functional regions
Group Sparse inverse covariance for multi-subject connectome
Producing single subject maps of seed-to-voxel correlation
Regions extraction using dictionary learning and functional connectomes
Computing a Region of Interest (ROI) mask manually
Extracting signals from brain regions using the NiftiLabelsMasker
Regions Extraction of Default Mode Networks using Smith Atlas
Beta-Series Modeling for Task-Based Functional Connectivity and Decoding
Massively univariate analysis of a calculation task from the Localizer dataset
Massively univariate analysis of a motor task from the Localizer dataset
Massively univariate analysis of face vs house recognition
Multivariate decompositions: Independent component analysis of fMRI