Note
Go to the end to download the full example code. or to run this example in your browser via Binder
Basic nilearn example: manipulating and looking at data¶
A simple example showing how to load an existing Nifti file and use basic nilearn functionalities.
# Let us use a Nifti file that is shipped with nilearn
from nilearn.datasets import MNI152_FILE_PATH
# Note that the variable MNI152_FILE_PATH is just a path to a Nifti file
print(f"Path to MNI152 template: {MNI152_FILE_PATH!r}")
Path to MNI152 template: PosixPath('/home/runner/work/nilearn/nilearn/.tox/doc/lib/python3.10/site-packages/nilearn/datasets/data/mni_icbm152_t1_tal_nlin_sym_09a_converted.nii.gz')
A first step: looking at our data¶
Let’s quickly plot this file:
from nilearn import plotting
plotting.plot_img(MNI152_FILE_PATH)
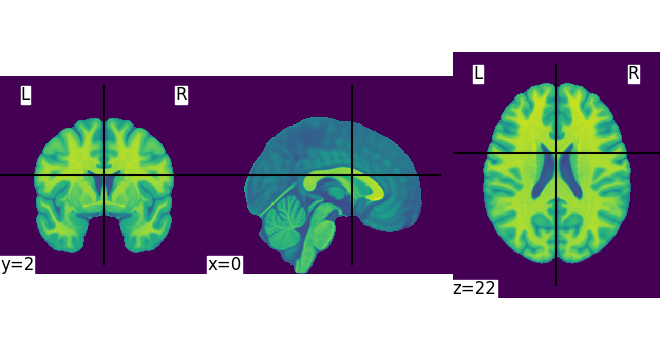
<nilearn.plotting.displays._slicers.OrthoSlicer object at 0x7f4a9d5a07c0>
This is not a very pretty plot. We just used the simplest possible code. There is a whole section of the documentation on making prettier code.
Exercise: Try plotting one of your own files. In the above, MNI152_FILE_PATH is nothing more than a string with a path pointing to a nifti image. You can replace it with a string pointing to a file on your disk. Note that it should be a 3D volume, and not a 4D volume.
Simple image manipulation: smoothing¶
Let’s use an image-smoothing function from nilearn:
smooth_img
Functions containing ‘img’ can take either a filename or an image as input.
Here we give as inputs the image filename and the smoothing value in mm
from nilearn import image
smooth_anat_img = image.smooth_img(MNI152_FILE_PATH, fwhm=3)
# While we are giving a file name as input, the function returns
# an in-memory object:
smooth_anat_img
<nibabel.nifti1.Nifti1Image object at 0x7f4a9d5a14e0>
This is an in-memory object. We can pass it to nilearn function, for instance to look at it
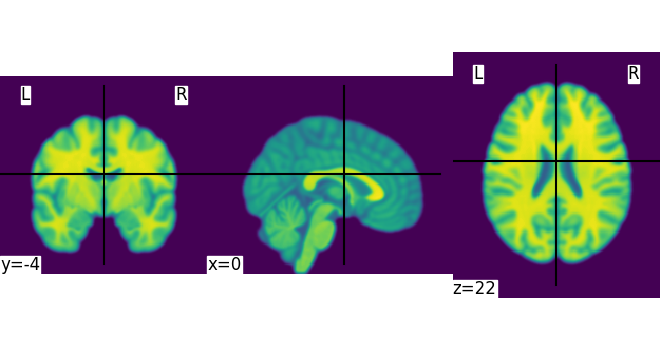
<nilearn.plotting.displays._slicers.OrthoSlicer object at 0x7f4a9dd60730>
We could also pass it to the smoothing function
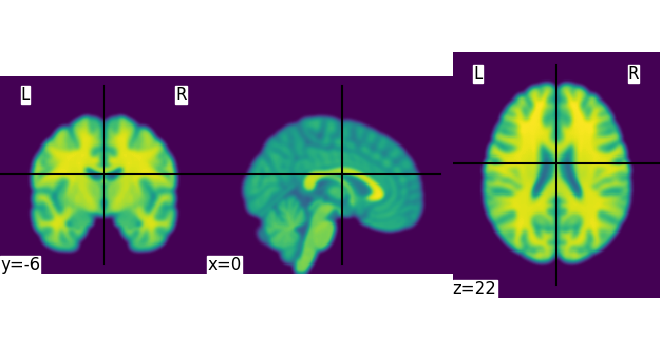
<nilearn.plotting.displays._slicers.OrthoSlicer object at 0x7f4a9d85b2e0>
Globbing over multiple 3D volumes¶
Nilearn also supports reading multiple volumes at once, using glob-style patterns. For instance, we can smooth volumes from many subjects at once and get a 4D image as output.
First let’s fetch Haxby dataset for subject 1 and 2
from nilearn import datasets
haxby = datasets.fetch_haxby(subjects=[1, 2])
[fetch_haxby] Dataset found in /home/runner/nilearn_data/haxby2001
[fetch_haxby] Downloading data from
http://data.pymvpa.org/datasets/haxby2001/subj1-2010.01.14.tar.gz ...
[fetch_haxby] Downloaded 183959552 of 314803244 bytes (58.4%%, 00 HR 00 MIN 01
SEC remaining)
[fetch_haxby] ...done. (2 seconds, 0 min)
[fetch_haxby] Extracting data from /home/runner/nilearn_data/haxby2001/b2fd65a88
d22090da62c3fb828be840e/subj1-2010.01.14.tar.gz...
[fetch_haxby] .. done.
Now we can find the anatomical images from both subjects using the * wildcard
from pathlib import Path
anats_all_subjects = (
Path(datasets.get_data_dirs()[0]) / "haxby2001" / "subj*" / "anat*"
)
Now we can smooth all the anatomical images at once
This is a 4D image containing one volume per subject
(124, 256, 256, 2)
Saving results to a file¶
We can save any in-memory object as follows:
output_dir = Path.cwd() / "results" / "plot_nilearn_101"
output_dir.mkdir(exist_ok=True, parents=True)
print(f"Output will be saved to: {output_dir}")
anats_all_subjects_smooth.to_filename(
output_dir / "anats_all_subjects_smooth.nii.gz"
)
Output will be saved to: /home/runner/work/nilearn/nilearn/examples/00_tutorials/results/plot_nilearn_101
Finally, calling plotting.show() is necessary to display the figure when running as a script outside IPython
To recap, all the nilearn tools can take data as filenames or glob-style patterns or in-memory objects, and return brain volumes as in-memory objects. These can be passed on to other nilearn tools, or saved to disk.
Total running time of the script: (0 minutes 7.938 seconds)
Estimated memory usage: 244 MB