Note
This page is a reference documentation. It only explains the function signature, and not how to use it. Please refer to the user guide for the big picture.
nilearn.masking.compute_epi_mask¶
- nilearn.masking.compute_epi_mask(epi_img, lower_cutoff=0.2, upper_cutoff=0.85, connected=True, opening=2, exclude_zeros=False, ensure_finite=True, target_affine=None, target_shape=None, memory=None, verbose=0)[source]¶
Compute a brain mask from fMRI data in 3D or 4D
numpy.ndarray.This is based on an heuristic proposed by T.Nichols: find the least dense point of the histogram, between fractions
lower_cutoffandupper_cutoffof the total image histogram.Note
In case of failure, it is usually advisable to increase
lower_cutoff.- Parameters:
- epi_imgNiimg-like object
See Input and output: neuroimaging data representation. EPI image, used to compute the mask. 3D and 4D images are accepted.
Note
If a 3D image is given, we suggest to use the mean image.
- lower_cutoff
float, optional Lower fraction of the histogram to be discarded. default=0.2.
- upper_cutoff
float, optional Upper fraction of the histogram to be discarded. default=0.85.
- connected
bool, optional If connected is True, only the largest connect component is kept. default=True.
- opening
boolorint, optional This parameter determines whether a morphological opening is performed, to keep only large structures. This step is useful to remove parts of the skull that might have been included. opening can be:
A
bool: If False, no opening is performed. If True, it is equivalent to opening=1.An
intn: The opening is performed via n erosions (seescipy.ndimage.binary_erosion). The largest connected component is then estimated if connected is set to True, and 2`n` dilation operations are performed (seescipy.ndimage.binary_dilation) followed by n erosions. This corresponds to 1 opening operation of order n followed by a closing operator of order n.
Note
Turning off opening (opening=False) will also prevent any smoothing applied to the image during the mask computation.
default=2.
- ensure_finite
bool, default=True If ensure_finite is True, the non-finite values (NaNs and infs) found in the images will be replaced by zeros
- exclude_zeros
bool, default=False Consider zeros as missing values for the computation of the threshold. This option is useful if the images have been resliced with a large padding of zeros.
- target_affine3x3 or a 4x4 array-like, or None, default=None
If specified, the image is resampled corresponding to this new affine.
Note
This parameter is passed to
nilearn.image.resample_img.- target_shape
tupleorlistor None, default=None If specified, the image will be resized to match this new shape. len(target_shape) must be equal to 3.
Note
If target_shape is specified, a target_affine of shape (4, 4) must also be given.
Note
This parameter is passed to
nilearn.image.resample_img.- memoryNone, instance of
joblib.Memory,str, orpathlib.Path, default=None Used to cache the masking process. By default, no caching is done. If a
stris given, it is the path to the caching directory.- verbose
boolorint, default=0 Verbosity level (
0orFalsemeans no message).
- Returns:
- mask
nibabel.nifti1.Nifti1Image The brain mask (3D image).
- mask
Examples using nilearn.masking.compute_epi_mask¶
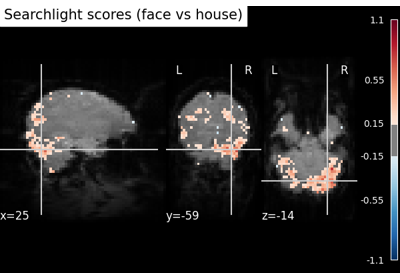
SearchLight MVPA with a custom correlation estimator on Haxby data