Note
This page is a reference documentation. It only explains the class signature, and not how to use it. Please refer to the user guide for the big picture.
nilearn.reporting.HTMLReport¶
- class nilearn.reporting.HTMLReport(head_tpl, body, head_values=None)[source]¶
A report written as HTML.
Methods such as
save_as_html, oropen_in_browserare inherited from classnilearn.plotting.html_document.HTMLDocument.- Parameters:
- head_tplstr.Template or Jinja Template
This is meant for display as a full page, like writing on disk. This is the Template object used to generate the HTML head section of the report. The template should be filled with:
title: The title of the HTML page.
- body: The full body of the HTML page. Provided through
the
bodyinput.
- body
str This parameter is used for embedding in the provided
head_tpltemplate. It contains the full body of the HTML page.- head_values
dict, default=None Additional substitutions in
head_tpl. ifNoneis passed, defaults to{}Note
This can be used to provide additional values with custom templates.
- get_iframe(width=None, height=None)[source]¶
Get the document wrapped in an inline frame.
For inserting in another HTML page of for display in a Jupyter notebook.
- property height¶
- open_in_browser(file_name=None)[source]¶
Save the plot to a temporary HTML file and open it in a browser.
- Parameters:
- file_name
strorNone, default=None HTML file to use as a temporary file.
- file_name
- save_as_html(file_name)[source]¶
Save the plot in an HTML file, that can later be opened in a browser.
- Parameters:
- file_name
str Path to the HTML file used for saving.
- file_name
- property width¶
Examples using nilearn.reporting.HTMLReport¶
Decoding of a dataset after GLM fit for signal extraction
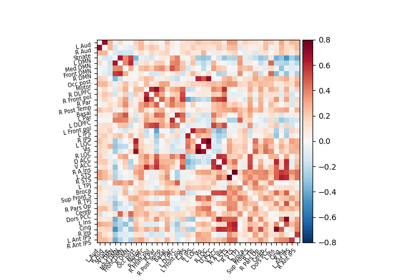
Extracting signals of a probabilistic atlas of functional regions
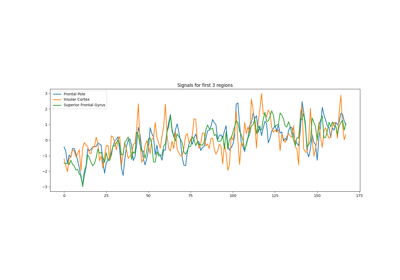
Extracting signals from brain regions using the NiftiLabelsMasker
Surface-based dataset first and second level analysis of a dataset