Note
Go to the end to download the full example code. or to run this example in your browser via Binder
Extracting signals from brain regions using the NiftiLabelsMasker¶
This simple example shows how to extract signals from functional
fMRI data and brain regions defined through an atlas.
More precisely, this example shows how to use the
NiftiLabelsMasker object to perform this
operation in just a few lines of code.
from nilearn._utils.helpers import check_matplotlib
check_matplotlib()
Retrieve the brain development functional dataset¶
We start by fetching the brain development functional dataset and we restrict the example to one subject only.
from nilearn.datasets import fetch_atlas_harvard_oxford, fetch_development_fmri
dataset = fetch_development_fmri(n_subjects=1)
func_filename = dataset.func[0]
# print basic information on the dataset
print(f"First functional nifti image (4D) is at: {func_filename}")
[fetch_development_fmri] Dataset found in
/home/runner/nilearn_data/development_fmri
[fetch_development_fmri] Dataset found in
/home/runner/nilearn_data/development_fmri/development_fmri
[fetch_development_fmri] Dataset found in
/home/runner/nilearn_data/development_fmri/development_fmri
First functional nifti image (4D) is at: /home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar123_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz
Load an atlas¶
We then load the Harvard-Oxford atlas to define the brain regions and the first label correspond to the background.
atlas = fetch_atlas_harvard_oxford("cort-maxprob-thr25-2mm")
print(f"The atlas contains {len(atlas.labels) - 1} non-overlapping regions")
[fetch_atlas_harvard_oxford] Dataset found in /home/runner/nilearn_data/fsl
The atlas contains 48 non-overlapping regions
Instantiate the mask and visualize atlas¶
Instantiate the masker with label image and label values
from nilearn.maskers import NiftiLabelsMasker
masker = NiftiLabelsMasker(atlas.maps, lut=atlas.lut, verbose=1)
Visualize the atlas¶
We need to call fit prior to generating the mask. We can then generate a report to visualize the atlas. Here we use the ‘brainsprite’ engine that gives an interactive vizualtion instead of the static one generated by the matplotlib engine.
Note
The generated report can be:
displayed in a Notebook,
opened in a browser using the
.open_in_browser()method,or saved to a file using the
.save_as_html(output_filepath)method.
masker.fit()
report = masker.generate_report(engine="brainsprite")
report
\[NiftiLabelsMasker.fit] Loading regions from <nibabel.nifti1.Nifti1Image object
at 0x7f4a8ca98ee0>
\[NiftiLabelsMasker.fit] Finished fit
/home/runner/work/nilearn/nilearn/.tox/doc/lib/python3.10/site-packages/numpy/core/fromnumeric.py:771: UserWarning:
Warning: 'partition' will ignore the 'mask' of the MaskedArray.
/home/runner/work/nilearn/nilearn/examples/06_manipulating_images/plot_nifti_labels_simple.py:68: UserWarning:
No image provided to fit in NiftiLabelsMasker. Plotting ROIs of label image on the MNI152Template for reporting.
Fitting the masker on data and generating a report¶
We can again generate a report, but this time, the provided functional image is displayed with the ROI of the atlas. The report also contains a summary table giving the region sizes in mm3.
\[NiftiLabelsMasker.fit] Loading regions from <nibabel.nifti1.Nifti1Image object
at 0x7f4a8ca98ee0>
\[NiftiLabelsMasker.fit] Resampling regions
\[NiftiLabelsMasker.fit] Finished fit
Process the data with the NiftiLablesMasker¶
In order to extract the signals, we need to call transform on the functional data.
signals = masker.transform(func_filename)
# signals is a 2D numpy array, (n_time_points x n_regions)
print(f"{signals.shape=}")
/home/runner/work/nilearn/nilearn/examples/06_manipulating_images/plot_nifti_labels_simple.py:90: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
\[NiftiLabelsMasker.wrapped] Loading data from '/home/runner/nilearn_data/develo
pment_fmri/development_fmri/sub-pixar123_task-pixar_space-MNI152NLin2009cAsym_de
sc-preproc_bold.nii.gz'
\[NiftiLabelsMasker.wrapped] Extracting region signals
\[NiftiLabelsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/06_manipulating_images/plot_nifti_labels_simple.py:90: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
signals.shape=(168, 48)
Output to dataframe and plot¶
You can use ‘set_output()’ to decide the output format of ‘transform’. If you want to output to a DataFrame, you an choose pandas and polars.
masker.set_output(transform="pandas")
signals_df = masker.transform(func_filename)
print(signals_df.head)
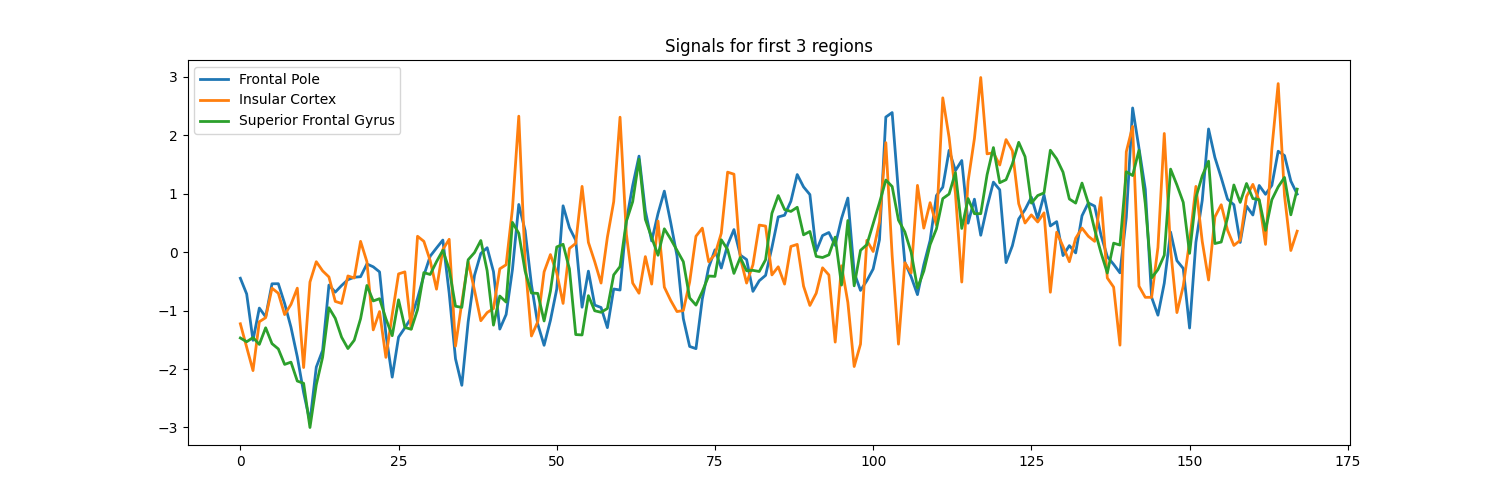
signals_df[["Frontal Pole", "Insular Cortex", "Superior Frontal Gyrus"]].plot(
title="Signals from 3 regions", figsize=(15, 5)
)
/home/runner/work/nilearn/nilearn/examples/06_manipulating_images/plot_nifti_labels_simple.py:103: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
\[NiftiLabelsMasker.wrapped] Loading data from '/home/runner/nilearn_data/develo
pment_fmri/development_fmri/sub-pixar123_task-pixar_space-MNI152NLin2009cAsym_de
sc-preproc_bold.nii.gz'
\[NiftiLabelsMasker.wrapped] Extracting region signals
\[NiftiLabelsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/06_manipulating_images/plot_nifti_labels_simple.py:103: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
<bound method NDFrame.head of Frontal Pole Insular Cortex ... Supracalcarine Cortex Occipital Pole
0 635.218164 555.308283 ... 559.291453 681.952471
1 634.725215 554.842059 ... 563.142865 680.775976
2 633.273421 554.375836 ... 566.609135 681.662737
3 634.283365 555.348824 ... 569.112553 679.924334
4 634.009838 555.429906 ... 568.149700 679.476564
.. ... ... ... ... ...
163 638.127763 558.794824 ... 573.156535 691.601486
164 639.197822 560.092141 ... 575.467382 689.388973
165 639.062562 557.862377 ... 578.933652 689.915762
166 638.263023 556.767765 ... 577.393087 688.098340
167 637.854236 557.152906 ... 572.193682 687.685689
[168 rows x 48 columns]>
<Axes: title={'center': 'Signals from 3 regions'}>
Total running time of the script: (0 minutes 6.760 seconds)
Estimated memory usage: 411 MB