Note
This page is a reference documentation. It only explains the class signature, and not how to use it. Please refer to the user guide for the big picture.
nilearn.glm.LikelihoodModelResults¶
- class nilearn.glm.LikelihoodModelResults(theta, Y, model, cov=None, dispersion=1.0, nuisance=None)[source]¶
Class to contain results from likelihood models.
This is the class in which things like AIC, BIC, llf can be implemented as methods, not computed in, say, the fit method of OLSModel.
- Parameters:
- thetandarray
Parameter estimates from estimated model.
- Yndarray
Data.
- model
LikelihoodModelinstance Model used to generate fit.
- covNone or ndarray, default=None
Covariance of thetas.
- dispersionscalar, default=1
Multiplicative factor in front of cov.
- nuisanceNone of ndarray, default=None
Parameter estimates needed to compute logL.
Notes
The covariance of thetas is given by:
dispersion * cov
For (some subset of models) dispersion will typically be the mean square error from the estimated model (sigma^2)
- Fcontrast(matrix, dispersion=None, invcov=None)[source]¶
Compute an F contrast for a contrast matrix
matrix.Here,
matrixM is assumed to be non-singular. More precisely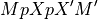
is assumed invertible. Here,
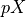 is the generalized inverse of
the design matrix of the model.
There can be problems in non-OLS models where
the rank of the covariance of the noise is not full.
is the generalized inverse of
the design matrix of the model.
There can be problems in non-OLS models where
the rank of the covariance of the noise is not full.See the contrasts module to see how to specify contrasts. In particular, the matrices from these contrasts will always be non-singular in the sense above.
- Parameters:
- matrix1D array-like
Contrast matrix.
- dispersionNone or
float, default=None If None, use
self.dispersion.- invcovNone or array, default=None
Known inverse of variance covariance matrix. If None, calculate this matrix.
- Returns:
- f_res
FContrastResultsinstance with attributes F, df_den, df_num
- f_res
Notes
For F contrasts, we now specify an effect and covariance.
- Tcontrast(matrix, store=('t', 'effect', 'sd'), dispersion=None)[source]¶
Compute a Tcontrast for a row vector matrix.
To get the t-statistic for a single column, use the ‘t’ method.
- Parameters:
- matrix1D array-like
Contrast matrix.
- storesequence, default=(‘t’, ‘effect’, ‘sd’)
Components of t to store in results output object.
- dispersionNone or
float, default = None
- Returns:
- res
TContrastResultsobject
- res
- conf_int(alpha=0.05, cols=None, dispersion=None)[source]¶
Return the confidence interval of the specified theta estimates.
- Parameters:
- alpha
float, default=0.05 The alpha level for the confidence interval. ie., alpha = .05 returns a 95% confidence interval.
- cols
tuple, default=None cols specifies which confidence intervals to return.
- dispersionNone or scalar, default=None
Scale factor for the variance / covariance (see class docstring and
vcovmethod docstring).
- alpha
- Returns:
- cisndarray
cis is shape
(len(cols), 2)where each row contains [lower, upper] for the given entry in cols
Notes
Confidence intervals are two-tailed.
Examples
>>> from numpy.random import standard_normal as stan >>> from nilearn.glm import OLSModel >>> x = np.hstack((stan((30, 1)), stan((30, 1)), stan((30, 1)))) >>> beta = np.array([3.25, 1.5, 7.0]) >>> y = np.dot(x, beta) + stan((30)) >>> model = OLSModel(x).fit(y) >>> confidence_intervals = model.conf_int(cols=(1, 2))
- t(column=None)[source]¶
Return the (Wald) t-statistic for a given parameter estimate.
Use Tcontrast for more complicated (Wald) t-statistics.
- vcov(matrix=None, column=None, dispersion=None, other=None)[source]¶
Return Variance/covariance matrix of linear contrast.
- Parameters:
- matrix(dim, self.theta.shape[0]) array, default=None
Numerical contrast specification, where
dimrefers to the ‘dimension’ of the contrast i.e. 1 for t contrasts, 1 or more for F contrasts.- column
int, default=None Alternative way of specifying contrasts (column index).
- dispersion
floator (n_voxels,) array, default=None Value(s) for the dispersion parameters.
- other(dim, self.theta.shape[0]) array, default=None
Alternative contrast specification (?).
- Returns:
- cov(dim, dim) or (n_voxels, dim, dim) array
The estimated covariance matrix/matrices.
- Returns the variance/covariance matrix of a linear contrast of the
- estimates of theta, multiplied by dispersion which will often be an
- estimate of dispersion, like, sigma^2.
- The covariance of interest is either specified as a (set of) column(s)
- or a matrix.