Note
This page is a reference documentation. It only explains the class signature, and not how to use it. Please refer to the user guide for the big picture.
nilearn.decomposition.DictLearning¶
- class nilearn.decomposition.DictLearning(n_components=20, n_epochs=1, alpha=10, reduction_ratio='auto', dict_init=None, random_state=None, batch_size=20, method='cd', mask=None, smoothing_fwhm=4, standardize=True, standardize_confounds=True, detrend=True, low_pass=None, high_pass=None, t_r=None, target_affine=None, target_shape=None, mask_strategy='epi', mask_args=None, n_jobs=1, verbose=0, memory=None, memory_level=0)[source]¶
Perform a map learning algorithm based on spatial component sparsity, over a CanICA initialization.
This yields more stable maps than CanICA.
See Mensch et al.[1].
Added in Nilearn 0.2.
- Parameters:
- n_components
int, default=20 Number of components to extract.
- n_epochs
float, default=1 Number of epochs the algorithm should run on the data.
- alpha
float, default=10 Sparsity controlling parameter.
- reduction_ratio‘auto’ or
floatbetween 0. and 1., default=’auto’ Between 0. or 1. : controls data reduction in the temporal domain. 1. means no reduction, < 1. calls for an SVD based reduction.
if set to ‘auto’, estimator will set the number of components per reduced session to be n_components.
- dict_initNiimg-like object or
SurfaceImageor None, default=None Initial estimation of dictionary maps. Would be computed from CanICA if not provided.
- random_state
intor np.random.RandomState, optional Pseudo-random number generator state used for random sampling.
- batch_size
int, default=20 The number of samples to take in each batch.
- method{‘cd’, ‘lars’}, default=’cd’
Coding method used by sklearn backend. Below are the possible values. lars: uses the least angle regression method to solve the lasso problem (linear_model.lars_path) cd: uses the coordinate descent method to compute the Lasso solution (linear_model.Lasso). Lars will be faster if the estimated components are sparse.
- maskNiimg-like object,
MultiNiftiMaskerorSurfaceImageorMultiSurfaceMaskerobject, or None default=None Mask to be used on data. If an instance of masker is passed, then its mask will be used. If no mask is given, for Nifti images, it will be computed automatically by a MultiNiftiMasker with default parameters; for surface images, all the vertices will be used.
- smoothing_fwhm
floatorintor None, optional. If smoothing_fwhm is not None, it gives the full-width at half maximum in millimeters of the spatial smoothing to apply to the signal. default=4mm.
- standardizeany of: ‘zscore_sample’, ‘zscore’, ‘psc’, True, False or None; default=True
Strategy to standardize the signal:
'zscore_sample': The signal is z-scored. Timeseries are shifted to zero mean and scaled to unit variance. Uses sample std.'psc': Timeseries are shifted to zero mean value and scaled to percent signal change (as compared to original mean signal).True: The signal is z-scored (same as option zscore). Timeseries are shifted to zero mean and scaled to unit variance.Deprecated since Nilearn 0.13.0: In nilearn version 0.15.0,
Truewill be replaced by'zscore_sample'.False: Do not standardize the data.Deprecated since Nilearn 0.13.0: In nilearn version 0.15.0,
Falsewill be replaced byNone.
Deprecated since Nilearn 0.13.0: The default will be changed to
'zscore_sample'in version 0.15.0.- standardize_confounds
bool, default=True If set to True, the confounds are z-scored: their mean is put to 0 and their variance to 1 in the time dimension.
- detrend
bool, optional Whether to detrend signals or not.
- low_pass
floatorintor None, default=None Low cutoff frequency in Hertz. If specified, signals above this frequency will be filtered out. If None, no low-pass filtering will be performed.
Note
This parameter is passed to
nilearn.image.resample_img.- high_pass
floatorintor None, default=None High cutoff frequency in Hertz. If specified, signals below this frequency will be filtered out.
Note
This parameter is passed to
nilearn.image.resample_img.- t_r
floatorintor None, default=None Repetition time, in seconds (sampling period). Set to None if not provided.
Note
This parameter is passed to
nilearn.image.resample_img.- target_affine3x3 or a 4x4 array-like, or None, default=None
If specified, the image is resampled corresponding to this new affine.
Note
This parameter is passed to
nilearn.image.resample_img.- target_shape
tupleorlistor None, default=None If specified, the image will be resized to match this new shape. len(target_shape) must be equal to 3.
Note
If target_shape is specified, a target_affine of shape (4, 4) must also be given.
Note
This parameter is passed to
nilearn.image.resample_img.- mask_strategy{“background”, “epi”, “whole-brain-template”,”gm-template”, “wm-template”}, optional
The strategy used to compute the mask:
"background": Use this option if your images present a clear homogeneous background. Usesnilearn.masking.compute_background_maskunder the hood."epi": Use this option if your images are raw EPI images. Usesnilearn.masking.compute_epi_mask."whole-brain-template": This will extract the whole-brain part of your data by resampling the MNI152 brain mask for your data’s field of view. Usesnilearn.masking.compute_brain_maskwithmask_type="whole-brain".Note
This option is equivalent to the previous ‘template’ option which is now deprecated.
"gm-template": This will extract the gray matter part of your data by resampling the corresponding MNI152 template for your data’s field of view. Usesnilearn.masking.compute_brain_maskwithmask_type="gm".Added in Nilearn 0.8.1.
"wm-template": This will extract the white matter part of your data by resampling the corresponding MNI152 template for your data’s field of view. Usesnilearn.masking.compute_brain_maskwithmask_type="wm".Added in Nilearn 0.8.1.
default=’epi’.
Note
These strategies are only relevant for Nifti images and the parameter is ignored for SurfaceImage objects.
- mask_args
dictor None, default=None If mask is None, these are additional parameters passed to
nilearn.masking.compute_background_mask, ornilearn.masking.compute_epi_maskto fine-tune mask computation. Please see the related documentation for details.- n_jobs
int, default=1 The number of CPUs to use to do the computation. -1 means ‘all CPUs’.
- verbose
boolorint, default=0 Verbosity level (
0orFalsemeans no message).- memoryNone, instance of
joblib.Memory,str, orpathlib.Path, default=None Used to cache the masking process. By default, no caching is done. If a
stris given, it is the path to the caching directory.- memory_level
int, default=0 Rough estimator of the amount of memory used by caching. Higher value means more memory for caching. Zero means no caching.
- n_components
- Attributes:
- maps_masker_instance of NiftiMapsMasker or SurfaceMapsMasker
This masker was initialized with
components_img_,masker_.mask_img_and is the masker used when calling transform and inverse_transform.- mask_img_Niimg-like object or
SurfaceImage See Input and output: neuroimaging data representation. The mask of the data. If no mask was given at masker creation :
- for Nifti images, this contains automatically computed mask
via the selected
mask_strategy.
- for SurfaceImage objects, this mask encompasses all vertices of
the input images.
- components_2D numpy array (n_components x n-voxels or n-vertices)
Array of masked extracted components.
Note
Use attribute
components_img_rather than manually unmaskingcomponents_withmasker_attribute.- components_img_4D Nifti image or 2D
SurfaceImage The image giving the extracted components. Each 3D Nifti image or 1D SurfaceImage is a component.
Added in Nilearn 0.4.1.
- masker_
MultiNiftiMaskerorMultiSurfaceMasker Masker used to filter and mask data as first step. If
MultiNiftiMaskerorMultiSurfaceMaskeris given inmaskparameter, this is a copy of it. Otherwise, a masker is created using the value ofmaskand other Masker related parameters as initialization.- memory_joblib memory cache
- components_init_2D numpy array (n_components x n-voxels or n-vertices)
Array of components used for initialization.
- loadings_init_2D numpy array
Initial loadings.
References
- __init__(n_components=20, n_epochs=1, alpha=10, reduction_ratio='auto', dict_init=None, random_state=None, batch_size=20, method='cd', mask=None, smoothing_fwhm=4, standardize=True, standardize_confounds=True, detrend=True, low_pass=None, high_pass=None, t_r=None, target_affine=None, target_shape=None, mask_strategy='epi', mask_args=None, n_jobs=1, verbose=0, memory=None, memory_level=0)[source]¶
- fit(imgs, y=None, confounds=None)[source]¶
Compute the mask and the components across subjects.
- Parameters:
- imgslist of Niimg-like objects or list of
SurfaceImage See Input and output: neuroimaging data representation. Data on which the mask is calculated. If this is a list, the affine (for Niimg-like objects) and mesh (for SurfaceImages) is considered the same for all
- yNone
This parameter is unused. It is solely included for scikit-learn compatibility.
- confoundslist of CSV file paths, numpy.ndarrays or pandas DataFrames or None, default=None.
This parameter is passed to nilearn.signal.clean. Please see the related documentation for details. Should match with the list of imgs given.
- imgslist of Niimg-like objects or list of
- Returns:
- selfobject
Returns the instance itself. Contains attributes listed at the object level.
- fit_transform(X, y=None, **fit_params)¶
Fit to data, then transform it.
Fits transformer to X and y with optional parameters fit_params and returns a transformed version of X.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
Input samples.
- yarray-like of shape (n_samples,) or (n_samples, n_outputs), default=None
Target values (None for unsupervised transformations).
- **fit_paramsdict
Additional fit parameters.
- Returns:
- X_newndarray array of shape (n_samples, n_features_new)
Transformed array.
- get_metadata_routing()¶
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)¶
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- inverse_transform(loadings)[source]¶
Use provided loadings to compute corresponding linear component combination in whole-brain voxel space.
- Parameters:
- loadingslist of numpy array (n_samples x n_components)
Component signals to transform back into voxel signals
- Returns:
- reconstructed_imgslist of nibabel.Nifti1Image or
SurfaceImage - For each loading, reconstructed Nifti1Image or SurfaceImage.
- reconstructed_imgslist of nibabel.Nifti1Image or
- property n_elements_¶
- score(imgs, y=None, confounds=None, per_component=False)[source]¶
Score function based on explained variance on imgs.
Should only be used by DecompositionEstimator derived classes
- Parameters:
- imgsiterable of Niimg-like objects or
listofSurfaceImage See Input and output: neuroimaging data representation. Data to be scored
- %(y_dummy)s
- confoundsCSV file path or numpy.ndarray or pandas DataFrame or None, default=None
This parameter is passed to nilearn.signal.clean. Please see the related documentation for details
- per_componentbool, default=False
Specify whether the explained variance ratio is desired for each map or for the global set of components.
- imgsiterable of Niimg-like objects or
- Returns:
- scorefloat
Holds the score for each subjects. Score is two dimensional if per_component is True. First dimension is squeezed if the number of subjects is one
- set_fit_request(*, confounds='$UNCHANGED$', imgs='$UNCHANGED$')¶
Request metadata passed to the
fitmethod.Note that this method is only relevant if
enable_metadata_routing=True(seesklearn.set_config). Please see User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed tofitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it tofit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
Note
This method is only relevant if this estimator is used as a sub-estimator of a meta-estimator, e.g. used inside a
Pipeline. Otherwise it has no effect.- Parameters:
- confoundsstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
confoundsparameter infit.- imgsstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
imgsparameter infit.
- Returns:
- selfobject
The updated object.
- set_inverse_transform_request(*, loadings='$UNCHANGED$')¶
Request metadata passed to the
inverse_transformmethod.Note that this method is only relevant if
enable_metadata_routing=True(seesklearn.set_config). Please see User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed toinverse_transformif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it toinverse_transform.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
Note
This method is only relevant if this estimator is used as a sub-estimator of a meta-estimator, e.g. used inside a
Pipeline. Otherwise it has no effect.- Parameters:
- loadingsstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
loadingsparameter ininverse_transform.
- Returns:
- selfobject
The updated object.
- set_output(*, transform=None)[source]¶
Set the output container when
"transform"is called.Warning
This has not been implemented yet.
- set_params(**params)¶
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- set_score_request(*, confounds='$UNCHANGED$', imgs='$UNCHANGED$', per_component='$UNCHANGED$')¶
Request metadata passed to the
scoremethod.Note that this method is only relevant if
enable_metadata_routing=True(seesklearn.set_config). Please see User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed toscoreif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it toscore.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
Note
This method is only relevant if this estimator is used as a sub-estimator of a meta-estimator, e.g. used inside a
Pipeline. Otherwise it has no effect.- Parameters:
- confoundsstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
confoundsparameter inscore.- imgsstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
imgsparameter inscore.- per_componentstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
per_componentparameter inscore.
- Returns:
- selfobject
The updated object.
- set_transform_request(*, confounds='$UNCHANGED$', imgs='$UNCHANGED$')¶
Request metadata passed to the
transformmethod.Note that this method is only relevant if
enable_metadata_routing=True(seesklearn.set_config). Please see User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed totransformif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it totransform.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
Note
This method is only relevant if this estimator is used as a sub-estimator of a meta-estimator, e.g. used inside a
Pipeline. Otherwise it has no effect.- Parameters:
- confoundsstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
confoundsparameter intransform.- imgsstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
imgsparameter intransform.
- Returns:
- selfobject
The updated object.
- transform(imgs, confounds=None)[source]¶
Project the data into a reduced representation.
- Parameters:
- imgsiterable of Niimg-like objects or
listofSurfaceImage See Input and output: neuroimaging data representation. Data to be projected
- confoundsCSV file path or numpy.ndarray or pandas DataFrame or None, default=None
This parameter is passed to nilearn.signal.clean. Please see the related documentation for details
- imgsiterable of Niimg-like objects or
- Returns:
- loadingslist of 2D ndarray,
For each subject, each sample, loadings for each decomposition components shape: number of subjects * (number of scans, number of regions)
Examples using nilearn.decomposition.DictLearning¶
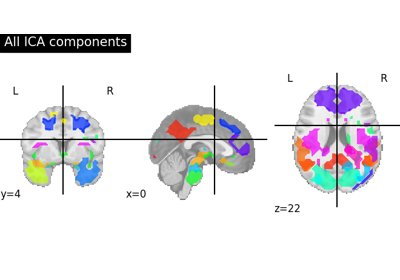
Deriving spatial maps from group fMRI data using ICA and Dictionary Learning
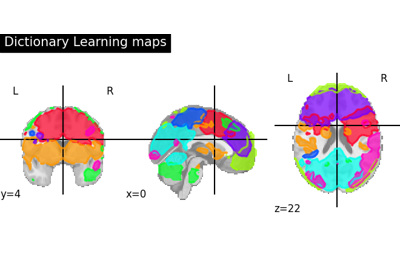
Regions extraction using dictionary learning and functional connectomes