Note
Go to the end to download the full example code. or to run this example in your browser via Binder
Making a surface plot of a 3D statistical map¶
Warning
This is an adaption of Making a surface plot of a 3D statistical map to use make it work with the new experimental surface API.
In this example, we will project a 3D statistical map onto a cortical mesh
using vol_to_surf,
display a surface plot of the projected map
using plot_surf_stat_map
with different plotting engines,
and add contours of regions of interest using
plot_surf_contours.
Get a statistical map¶
from nilearn import datasets
stat_img = datasets.load_sample_motor_activation_image()
Get a cortical mesh¶
from nilearn.experimental.surface import load_fsaverage, load_fsaverage_data
fsaverage_meshes = load_fsaverage()
Use mesh curvature to display useful anatomical information on inflated meshes
Here, we load the curvature map of the hemisphere under study, and define a surface map whose value for a given vertex is 1 if the curvature is positive, -1 if the curvature is negative.
import numpy as np
fsaverage_curvature = load_fsaverage_data(data_type="curvature")
curv_right_sign = np.sign(fsaverage_curvature.data.parts["right"])
Sample the 3D data around each node of the mesh¶
from nilearn.experimental.surface import SurfaceImage
img = SurfaceImage.from_volume(
mesh=fsaverage_meshes["pial"],
volume_img=stat_img,
)
Plot the result¶
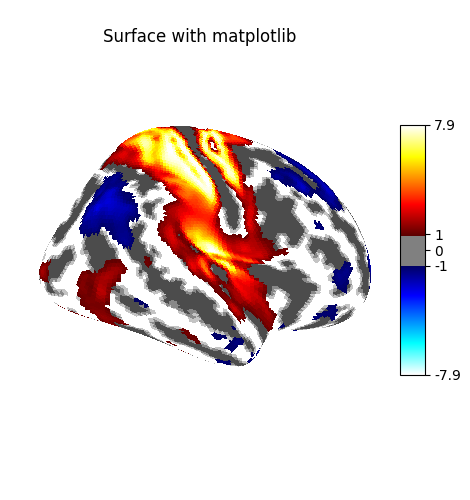
You can visualize the texture on the surface using the function
plot_surf_stat_map which uses matplotlib
as the default plotting engine.
from nilearn.experimental.plotting import plot_surf_stat_map
fig = plot_surf_stat_map(
stat_map=img,
surf_mesh=fsaverage_meshes["inflated"],
hemi="right",
title="Surface with matplotlib",
colorbar=True,
threshold=1.0,
bg_map=curv_right_sign,
)
fig.show()
Interactive plotting with Plotly¶
If you have a recent version of Nilearn (>=0.8.2), and if you have
plotly installed, you can easily configure
plot_surf_stat_map to use plotly
instead of matplotlib:
engine = "plotly"
# If plotly is not installed, use matplotlib
from nilearn._utils.helpers import is_plotly_installed
if not is_plotly_installed():
engine = "matplotlib"
print(f"Using plotting engine {engine}.")
fig = plot_surf_stat_map(
stat_map=img,
surf_mesh=fsaverage_meshes["inflated"],
hemi="right",
title="Surface with plotly",
colorbar=True,
threshold=1.0,
bg_map=curv_right_sign,
bg_on_data=True,
engine=engine, # Specify the plotting engine here
)
# Display the figure as with matplotlib figures
# fig.show()
Using plotting engine plotly.
/home/runner/work/nilearn/nilearn/.tox/doc/lib/python3.9/site-packages/nilearn/plotting/surf_plotting.py:983: UserWarning:
vmin cannot be chosen when cmap is symmetric
When using matplolib as the plotting engine, a standard
matplotlib.figure.Figure is returned.
With plotly as the plotting engine,
a custom PlotlySurfaceFigure
is returned which provides a similar API
to the Figure.
For example, you can save a static version of the figure to file
(this option requires to have kaleido installed):
# Save the figure as we would do with a matplotlib figure.
# Uncomment the following line to save the previous figure to file
# fig.savefig("right_hemisphere.png")
Plot 3D image for comparison¶
from nilearn import plotting
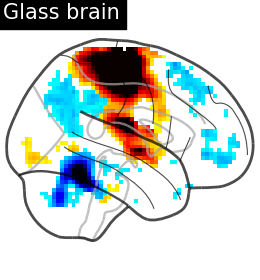
plotting.plot_glass_brain(
stat_map_img=stat_img,
display_mode="r",
plot_abs=False,
title="Glass brain",
threshold=2.0,
)
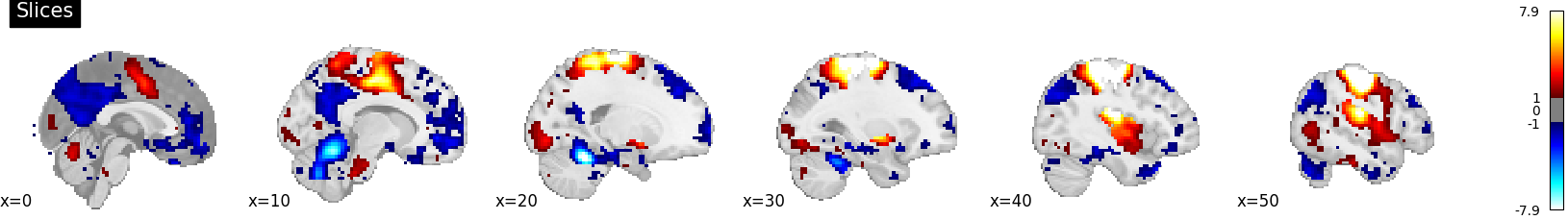
plotting.plot_stat_map(
stat_map_img=stat_img,
display_mode="x",
threshold=1.0,
cut_coords=range(0, 51, 10),
title="Slices",
)
<nilearn.plotting.displays._slicers.XSlicer object at 0x7f580d0c6070>
Use an atlas and choose regions to outline¶
from nilearn.experimental.surface import fetch_destrieux
destrieux_atlas, label_names = fetch_destrieux(mesh_type="inflated")
# these are the regions we want to outline
regions_dict = {
"G_postcentral": "Postcentral gyrus",
"G_precentral": "Precentral gyrus",
}
# get indices in atlas for these labels
regions_indices = [
np.where(np.array(label_names) == region)[0][0] for region in regions_dict
]
labels = list(regions_dict.values())
[get_dataset_dir] Dataset found in /home/runner/nilearn_data/destrieux_surface
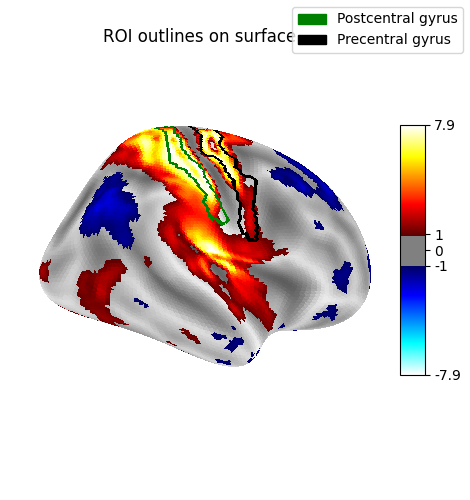
Display outlines of the regions of interest on top of a statistical map¶
from nilearn.experimental.plotting import plot_surf_contours
fsaverage_sulcal = load_fsaverage_data(data_type="sulcal", mesh_type="pial")
figure = plot_surf_stat_map(
stat_map=img,
surf_mesh=fsaverage_meshes["inflated"],
hemi="right",
title="ROI outlines on surface",
colorbar=True,
threshold=1.0,
bg_map=fsaverage_sulcal,
)
plot_surf_contours(
roi_map=destrieux_atlas,
hemi="right",
labels=labels,
levels=regions_indices,
figure=figure,
legend=True,
colors=["g", "k"],
)
plotting.show()
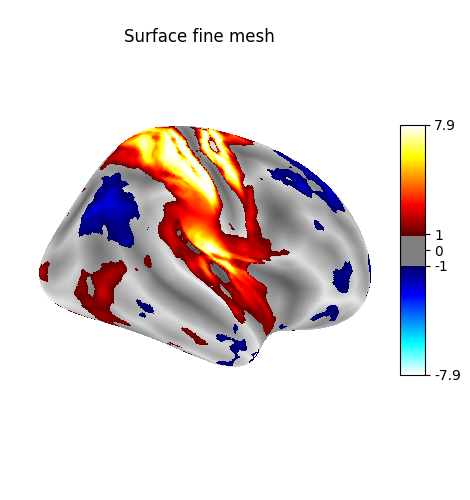
Plot with higher-resolution mesh¶
fetch_surf_fsaverage takes a mesh argument
which specifies whether to fetch the low-resolution fsaverage5 mesh,
or the high-resolution fsaverage mesh.
Using mesh="fsaverage" will result
in more memory usage and computation time, but finer visualizations.
big_fsaverage_meshes = load_fsaverage("fsaverage")
big_fsaverage_sulcal = load_fsaverage_data(
mesh_name="fsaverage", data_type="sulcal", mesh_type="inflated"
)
big_img = SurfaceImage.from_volume(
mesh=big_fsaverage_meshes["pial"],
volume_img=stat_img,
)
plot_surf_stat_map(
big_img,
surf_mesh=big_fsaverage_meshes["inflated"],
hemi="right",
colorbar=True,
title="Surface fine mesh",
threshold=1.0,
bg_map=big_fsaverage_sulcal,
)
[get_dataset_dir] Dataset found in /home/runner/nilearn_data/fsaverage
[get_dataset_dir] Dataset found in /home/runner/nilearn_data/fsaverage
[get_dataset_dir] Dataset found in /home/runner/nilearn_data/fsaverage
<Figure size 470x500 with 2 Axes>
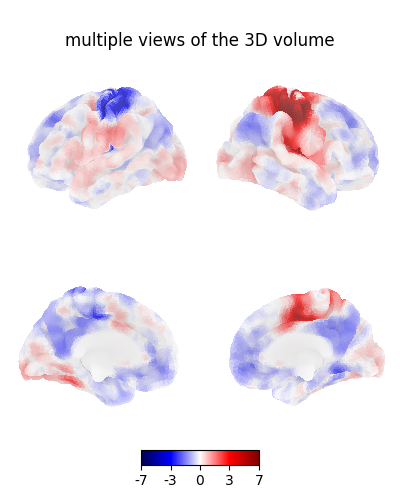
Plot multiple views of the 3D volume on a surface¶
plot_img_on_surf takes a statistical map
and projects it onto a surface.
It supports multiple choices of orientations,
and can plot either one or both hemispheres.
If no surf_mesh is given,
plot_img_on_surf projects the images onto
FreeSurfer's fsaverage5.
plotting.plot_img_on_surf(
stat_img,
views=["lateral", "medial"],
hemispheres=["left", "right"],
colorbar=True,
cmap="seismic",
title="multiple views of the 3D volume",
bg_on_data=True,
)
plotting.show()
3D visualization in a web browser¶
An alternative to nilearn.plotting.plot_surf_stat_map is to use
nilearn.plotting.view_surf or
nilearn.plotting.view_img_on_surf that give
more interactive visualizations in a web browser.
See 3D Plots of statistical maps or atlases on the cortical surface for more details.
from nilearn.experimental.plotting import view_surf
view = view_surf(
surf_mesh=fsaverage_meshes["inflated"],
surf_map=img,
threshold="90%",
bg_map=fsaverage_sulcal,
hemi="right",
title="3D visualization in a web browser",
)
# In a Jupyter notebook, if ``view`` is the output of a cell,
# it will be displayed below the cell
view
# view.open_in_browser()
# We don't need to do the projection ourselves, we can use
# :func:`~nilearn.plotting.view_img_on_surf`:
view = plotting.view_img_on_surf(stat_img, threshold="90%")
view
# view.open_in_browser()
Impact of plot parameters on visualization¶
You can specify arguments to be passed on to the function
nilearn.surface.vol_to_surf using vol_to_surf_kwargs
This allows fine-grained control of how the input 3D image
is resampled and interpolated -
for example if you are viewing a volumetric atlas,
you would want to avoid averaging the labels between neighboring regions.
Using nearest-neighbor interpolation with zero radius will achieve this.
destrieux = datasets.fetch_atlas_destrieux_2009(legacy_format=False)
view = plotting.view_img_on_surf(
destrieux.maps,
surf_mesh="fsaverage",
cmap="tab20",
vol_to_surf_kwargs={
"n_samples": 1,
"radius": 0.0,
"interpolation": "nearest",
},
symmetric_cmap=False,
colorbar=False,
)
view
# view.open_in_browser()
[get_dataset_dir] Dataset found in /home/runner/nilearn_data/destrieux_2009
[get_dataset_dir] Dataset found in /home/runner/nilearn_data/fsaverage
Total running time of the script: (0 minutes 26.165 seconds)
Estimated memory usage: 568 MB