Note
This page is a reference documentation. It only explains the class signature, and not how to use it. Please refer to the user guide for the big picture.
nilearn.decoding.SearchLight¶
- class nilearn.decoding.SearchLight(mask_img=None, process_mask_img=None, radius=2.0, estimator='svc', n_jobs=1, scoring=None, cv=None, verbose=0, random_state=0, estimator_args=None)[source]¶
Implement search_light analysis using an arbitrary type of classifier.
- Parameters:
- mask_imgNiimg-like object or None,
See Input and output: neuroimaging data representation. Boolean image giving location of voxels containing usable signals.
- process_mask_imgNiimg-like object or None, default=None
See Input and output: neuroimaging data representation. Boolean image giving voxels on which searchlight should be computed.
- radius
float, default=2. radius of the searchlight ball, in millimeters.
- estimator‘svr’, ‘svc’, or an estimator object implementing ‘fit’
The object to use to fit the data
- n_jobs
int, default=1 The number of CPUs to use to do the computation. -1 means ‘all CPUs’.
- scoring
stror callable or None, default=None The scoring strategy to use. See the scikit-learn documentation If callable, takes as arguments the fitted estimator, the test data (X_test) and the test target (y_test) if y is not None.
- cvcross-validation generator,
intor None, default=None A cross-validation generator. See: https://scikit-learn.org/stable/modules/cross_validation.html. If None is passed, cv=3 will be used. It can be an integer, in which case it is the number of folds in a KFold using
StratifiedKFoldwhen groups is None in thefitmethod for this class. If groups is specified butcvis not set to custom CV splitter, default isLeaveOneGroupOut.- verbose
boolorint, default=0 Verbosity level (
0orFalsemeans no message).- random_state
intor np.random.RandomState, optional Pseudo-random number generator state used for random sampling.
Note
This parameter will be passed to the underlying
"svc"estimator.- estimator_argsdict[str, Any] or None, default=None
Extra parameters to pass to the scikit-learn estimators.
Added in Nilearn 0.14.0dev.
- Attributes:
- mask_img_Nifti1Image or
SurfaceImage Mask computed by the masker object.
- masked_scores_numpy.ndarray
1D array containing the searchlight scores corresponding to the masked region only.
Added in Nilearn 0.11.0.
- n_elements_
int The number of voxels in the mask.
Added in Nilearn 0.12.1.
- process_mask_numpy.ndarray
- Boolean mask array representing the voxels included in the
searchlight computation.
Added in Nilearn 0.11.0.
- scores_numpy.ndarray
- 3D array containing searchlight scores for each voxel, aligned
with the mask.
Added in Nilearn 0.11.0.
- mask_img_Nifti1Image or
Notes
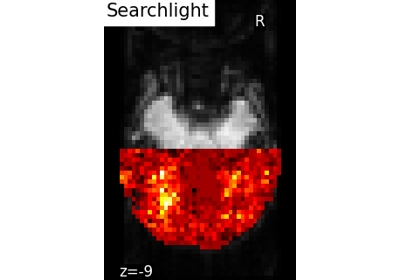
The searchlight [Kriegeskorte 06] is a widely used approach for the study of the fine-grained patterns of information in fMRI analysis. Its principle is relatively simple: a small group of neighboring features is extracted from the data, and the prediction function is instantiated on these features only. The resulting prediction accuracy is thus associated with all the features within the group, or only with the feature on the center. This yields a map of local fine-grained information, that can be used for assessing hypothesis on the local spatial layout of the neural code under investigation.
Nikolaus Kriegeskorte, Rainer Goebel & Peter Bandettini. Information-based functional brain mapping. Proceedings of the National Academy of Sciences of the United States of America, vol. 103, no. 10, pages 3863-3868, March 2006
- __init__(mask_img=None, process_mask_img=None, radius=2.0, estimator='svc', n_jobs=1, scoring=None, cv=None, verbose=0, random_state=0, estimator_args=None)[source]¶
- fit(imgs, y, groups=None)[source]¶
Fit the searchlight.
- Parameters:
- imgsNiimg-like object
See Input and output: neuroimaging data representation. 4D image.
- y1D array-like
Target variable to predict. Must have exactly as many elements as 3D images in img.
- groupsarray-like, default=None
group label for each sample for cross validation. Must have exactly as many elements as 3D images in img.
- fit_transform(imgs, y, groups=None)[source]¶
Fit the searchlight and applies to the input image.
- Parameters:
- imgsNiimg-like object
See Input and output: neuroimaging data representation. 4D image.
- y1D array-like
Target variable to predict. Must have exactly as many elements as 3D images in imgs.
- groupsarray-like, default=None
group label for each sample for cross validation. Must have exactly as many elements as 3D images in imgs.
- Returns:
- resultnp.ndarray
- get_metadata_routing()¶
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)¶
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- property scores_img_¶
Convert the 3D scores array into a NIfTI image.
- set_fit_request(*, groups='$UNCHANGED$', imgs='$UNCHANGED$')¶
Request metadata passed to the
fitmethod.Note that this method is only relevant if
enable_metadata_routing=True(seesklearn.set_config). Please see User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed tofitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it tofit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
Note
This method is only relevant if this estimator is used as a sub-estimator of a meta-estimator, e.g. used inside a
Pipeline. Otherwise it has no effect.- Parameters:
- groupsstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
groupsparameter infit.- imgsstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
imgsparameter infit.
- Returns:
- selfobject
The updated object.
- set_output(*, transform=None)[source]¶
Set the output container when
"transform"is called.Warning
This has not been implemented yet.
- set_params(**params)¶
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- set_transform_request(*, imgs='$UNCHANGED$')¶
Request metadata passed to the
transformmethod.Note that this method is only relevant if
enable_metadata_routing=True(seesklearn.set_config). Please see User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed totransformif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it totransform.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
Note
This method is only relevant if this estimator is used as a sub-estimator of a meta-estimator, e.g. used inside a
Pipeline. Otherwise it has no effect.- Parameters:
- imgsstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
imgsparameter intransform.
- Returns:
- selfobject
The updated object.
- transform(imgs)[source]¶
Apply the fitted searchlight on new images.
- Parameters:
- imgsNiimg-like object
See Input and output: neuroimaging data representation. 4D image.
- Returns:
- resultnp.ndarray