Note
Go to the end to download the full example code. or to run this example in your browser via Binder
Functional connectivity predicts age group¶
This example compares different kinds of functional connectivity between regions of interest : correlation, partial correlation, and tangent space embedding.
The resulting connectivity coefficients can be used to discriminate children from adults. In general, the tangent space embedding outperforms the standard correlations: see Dadi et al.[1] for a careful study.
try:
import matplotlib.pyplot as plt
except ImportError as e:
raise RuntimeError("This script needs the matplotlib library") from e
Load brain development fMRI dataset and MSDL atlas¶
We study only 60 subjects from the dataset, to save computation time.
from nilearn.datasets import fetch_atlas_msdl, fetch_development_fmri
development_dataset = fetch_development_fmri(n_subjects=60)
[fetch_development_fmri] Dataset found in
/home/runner/nilearn_data/development_fmri
[fetch_development_fmri] Dataset found in
/home/runner/nilearn_data/development_fmri/development_fmri
[fetch_development_fmri] Dataset found in
/home/runner/nilearn_data/development_fmri/development_fmri
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3f72286e80019c3c1af/ ...
[fetch_development_fmri] ...done. (4 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3f92286e80018c3e463/ ...
[fetch_development_fmri] ...done. (6 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3f6a743a90017608171/ ...
[fetch_development_fmri] ...done. (5 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3f64712b400183b70d8/ ...
[fetch_development_fmri] ...done. (6 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3eb2286e80019c3c198/ ...
[fetch_development_fmri] ...done. (4 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3ed2286e80017c41b56/ ...
[fetch_development_fmri] ...done. (6 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3ee2286e80016c3c379/ ...
[fetch_development_fmri] ...done. (4 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3ee4712b400183b70c3/ ...
[fetch_development_fmri] ...done. (5 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3efa743a9001660a0d5/ ...
[fetch_development_fmri] ...done. (5 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3f14712b4001a3b560e/ ...
[fetch_development_fmri] ...done. (5 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3f1a743a90017608164/ ...
[fetch_development_fmri] ...done. (6 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3f12286e80016c3c37e/ ...
[fetch_development_fmri] ...done. (5 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3f34712b4001a3b5612/ ...
[fetch_development_fmri] ...done. (3 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3f7a743a90019606cdf/ ...
[fetch_development_fmri] ...done. (4 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb470153992690018133d3b/ ...
[fetch_development_fmri] ...done. (5 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb46e793992690017108eb9/ ...
[fetch_development_fmri] ...done. (4 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb47038353c5800199ac9a2/ ...
[fetch_development_fmri] ...done. (5 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb46e85a3bc97001aeff750/ ...
[fetch_development_fmri] ...done. (6 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb4701c3992690018133d49/ ...
[fetch_development_fmri] ...done. (5 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb46e1c3992690018133a9e/ ...
[fetch_development_fmri] ...done. (7 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff39aa743a900176080bf/ ...
[fetch_development_fmri] ...done. (3 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff39d4712b400193b5b89/ ...
[fetch_development_fmri] ...done. (4 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb4703039926900160f6b3e/ ...
[fetch_development_fmri] ...done. (5 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb46e4d353c58001b9cb325/ ...
[fetch_development_fmri] ...done. (3 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb4700af2be3c0017056f69/ ...
[fetch_development_fmri] ...done. (5 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb46e0cf2be3c001801f757/ ...
[fetch_development_fmri] ...done. (4 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb4702b39926900171090e4/ ...
[fetch_development_fmri] ...done. (4 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb46e35f2be3c00190305ff/ ...
[fetch_development_fmri] ...done. (3 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3a12286e80017c41a48/ ...
[fetch_development_fmri] ...done. (3 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3a12286e80016c3c2fc/ ...
[fetch_development_fmri] ...done. (7 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb4701ff2be3c0017056fad/ ...
[fetch_development_fmri] ...done. (3 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb46e0339926900160f6930/ ...
[fetch_development_fmri] ...done. (6 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff39fa743a90018606e2f/ ...
[fetch_development_fmri] ...done. (6 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3a34712b4001a3b55a3/ ...
[fetch_development_fmri] ...done. (4 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb4703439926900160f6b43/ ...
[fetch_development_fmri] ...done. (3 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb46e40f2be3c001801f77f/ ...
[fetch_development_fmri] ...done. (5 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3a34712b400193b5b92/ ...
[fetch_development_fmri] ...done. (5 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3a84712b400183b7048/ ...
[fetch_development_fmri] ...done. (6 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff39aa743a90018606e21/ ...
[fetch_development_fmri] ...done. (6 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff39aa743a900176080ba/ ...
[fetch_development_fmri] ...done. (5 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3a72286e80017c41a54/ ...
[fetch_development_fmri] ...done. (6 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3a7a743a90018606e42/ ...
[fetch_development_fmri] ...done. (6 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb4702639926900190faf1d/ ...
[fetch_development_fmri] ...done. (4 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb46e3f353c5800199ac787/ ...
[fetch_development_fmri] ...done. (5 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff39ca743a90019606c50/ ...
[fetch_development_fmri] ...done. (3 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3a2a743a9001660a048/ ...
[fetch_development_fmri] ...done. (5 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb47020f2be3c0019030968/ ...
[fetch_development_fmri] ...done. (5 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb46e6f353c58001a9b3311/ ...
[fetch_development_fmri] ...done. (6 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb47016a3bc970018f1fc88/ ...
[fetch_development_fmri] ...done. (6 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb46e6ba3bc970019f07152/ ...
[fetch_development_fmri] ...done. (4 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff395a743a900176080af/ ...
[fetch_development_fmri] ...done. (6 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3964712b400193b5b7d/ ...
[fetch_development_fmri] ...done. (4 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb47057f2be3c0019030a1f/ ...
[fetch_development_fmri] ...done. (5 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb46e63f2be3c0017056ba9/ ...
[fetch_development_fmri] ...done. (4 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb4704af2be3c001705703b/ ...
[fetch_development_fmri] ...done. (4 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5cb46e7a353c58001a9b3324/ ...
[fetch_development_fmri] ...done. (4 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3952286e80016c3c2e7/ ...
[fetch_development_fmri] ...done. (3 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3954712b400193b5b79/ ...
[fetch_development_fmri] ...done. (4 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff399a743a9001660a031/ ...
[fetch_development_fmri] ...done. (4 seconds, 0 min)
[fetch_development_fmri] Downloading data from
https://osf.io/download/5c8ff3982286e80017c41a29/ ...
[fetch_development_fmri] ...done. (4 seconds, 0 min)
We use probabilistic regions of interest (ROIs) from the MSDL atlas.
from nilearn.maskers import NiftiMapsMasker
msdl_data = fetch_atlas_msdl()
msdl_coords = msdl_data.region_coords
masker = NiftiMapsMasker(
msdl_data.maps,
resampling_target="data",
t_r=development_dataset.t_r,
detrend=True,
low_pass=0.1,
high_pass=0.01,
memory="nilearn_cache",
memory_level=1,
standardize_confounds=True,
verbose=1,
)
masked_data = list(
map(
masker.fit_transform,
development_dataset.func,
development_dataset.confounds,
)
)
[fetch_atlas_msdl] Dataset found in /home/runner/nilearn_data/msdl_atlas
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar135_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe008d77fd0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar135_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.9s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar123_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe008d749a0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar123_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar124_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe008d77820>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar124_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar125_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe023472530>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar125_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar126_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe008d770d0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar126_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar127_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe008d77a30>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar127_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar134_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe008d77130>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar134_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar129_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe0234735b0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar129_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar130_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe008d75e10>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar130_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar131_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe02fd558d0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar131_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar132_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe023505f30>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar132_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar133_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe00a23c430>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar133_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar128_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe008d77a60>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar128_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar032_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe02fd55c00>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar032_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar033_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe02fd55c00>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar033_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar034_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe02fd558d0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar034_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar035_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fdfdfbc1cf0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar035_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar036_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe008d76440>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar036_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar037_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe008d75fc0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar037_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar038_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe009e16140>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar038_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar041_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fdfdfbc0940>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar041_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar040_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fdfc8b5e9e0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar040_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar042_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fdfc8b5ec80>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar042_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar043_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe023504ac0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar043_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar044_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe009e16950>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar044_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar031_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe009e160b0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar031_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar046_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe008d773d0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar046_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar047_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fdfdfbc0280>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar047_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar039_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe008d773d0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar039_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar045_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fdfc8b5c3d0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar045_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar001_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe0018db970>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar001_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar028_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fdfc8b5ff40>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar028_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.9s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar002_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe0018d9780>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar002_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar003_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe008d76500>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar003_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar004_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe023470280>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar004_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar005_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fdfc8b5d990>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar005_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar006_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe008d773d0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar006_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar007_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe008d76500>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar007_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar008_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe0018d8100>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar008_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar009_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fdfdfbc3c40>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar009_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar010_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe002dd4f10>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar010_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar011_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fdfdfbc1cf0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar011_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar012_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe0300d0400>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar012_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar013_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe0018d8a60>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar013_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar029_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe008d76440>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar029_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar014_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe023504ac0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar014_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar016_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe008d75b10>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar016_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar017_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe023471ff0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar017_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar018_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe009e160b0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar018_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar019_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fdfe201e710>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar019_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar020_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe008d76500>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar020_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar021_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fdfe201cf70>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar021_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar022_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe02db3f490>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar022_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar023_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe002dd6620>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar023_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar024_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe009e14520>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar024_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar025_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fdfdfbc1cf0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar025_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar026_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe009e160b0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar026_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar027_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fe0018d8a60>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar027_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar015_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fdfe201dbd0>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar015_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
________________________________________________________________________________
[Memory] Calling nilearn.maskers.base_masker.filter_and_extract...
filter_and_extract('/home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar030_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz',
<nilearn.maskers.nifti_maps_masker._ExtractionFunctor object at 0x7fdfe201cf70>, { 'allow_overlap': True,
'clean_args': None,
'clean_kwargs': {},
'cmap': 'CMRmap_r',
'detrend': True,
'dtype': None,
'high_pass': 0.01,
'high_variance_confounds': False,
'keep_masked_maps': False,
'low_pass': 0.1,
'maps_img': '/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii',
'mask_img': None,
'reports': True,
'smoothing_fwhm': None,
'standardize': False,
'standardize_confounds': True,
't_r': 2,
'target_affine': None,
'target_shape': None}, confounds=None, sample_mask=None, dtype=None, memory=Memory(location=nilearn_cache/joblib), memory_level=1, verbose=1, sklearn_output_config=None)
\[NiftiMapsMasker.wrapped] Loading data from '/home/runner/nilearn_data/developm
ent_fmri/development_fmri/sub-pixar030_task-pixar_space-MNI152NLin2009cAsym_desc
-preproc_bold.nii.gz'
\[NiftiMapsMasker.wrapped] Extracting region signals
\[NiftiMapsMasker.wrapped] Cleaning extracted signals
/home/runner/work/nilearn/nilearn/examples/07_advanced/plot_age_group_prediction_cross_val.py:50: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
_______________________________________________filter_and_extract - 0.8s, 0.0min
What kind of connectivity is most powerful for classification?¶
we will use connectivity matrices as features to distinguish children from adults. We use cross-validation and measure classification accuracy to compare the different kinds of connectivity matrices.
# prepare the classification pipeline
from sklearn.dummy import DummyClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import Pipeline
from sklearn.svm import LinearSVC
from nilearn.connectome import ConnectivityMeasure
kinds = ["correlation", "partial correlation", "tangent"]
pipe = Pipeline(
[
(
"connectivity",
ConnectivityMeasure(
vectorize=True,
),
),
(
"classifier",
GridSearchCV(
LinearSVC(dual=True, random_state=0),
{"C": [0.1, 1.0, 10.0]},
cv=5,
),
),
]
)
param_grid = [
{"classifier": [DummyClassifier(strategy="most_frequent")]},
{"connectivity__kind": kinds},
]
We use random splits of the subjects into training/testing sets. StratifiedShuffleSplit allows preserving the proportion of children in the test set.
from sklearn.model_selection import StratifiedShuffleSplit
from sklearn.preprocessing import LabelEncoder
groups = development_dataset.phenotypic["Child_Adult"].to_list()
classes = LabelEncoder().fit_transform(groups)
cv = StratifiedShuffleSplit(n_splits=30, random_state=0, test_size=10)
gs = GridSearchCV(
pipe,
param_grid,
scoring="accuracy",
cv=cv,
refit=False,
n_jobs=2,
)
gs.fit(masked_data, classes)
mean_scores = gs.cv_results_["mean_test_score"]
scores_std = gs.cv_results_["std_test_score"]
display the results
plt.figure(figsize=(6, 4), constrained_layout=True)
positions = [0.1, 0.2, 0.3, 0.4]
plt.barh(positions, mean_scores, align="center", height=0.05, xerr=scores_std)
yticks = ["dummy", *list(gs.cv_results_["param_connectivity__kind"].data[1:])]
yticks = [t.replace(" ", "\n") for t in yticks]
plt.yticks(positions, yticks)
plt.xlabel("Classification accuracy")
plt.gca().grid(True)
plt.gca().set_axisbelow(True)
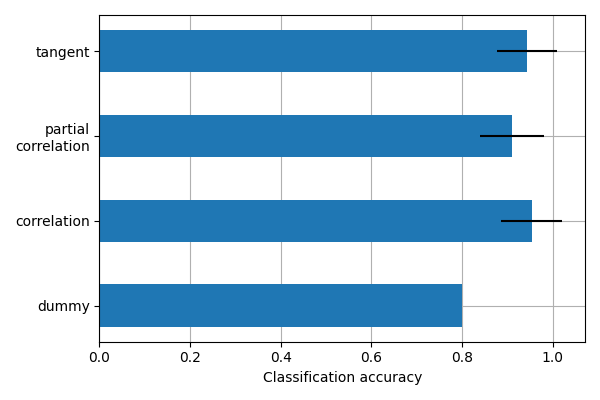
This is a small example to showcase nilearn features. In practice such comparisons need to be performed on much larger cohorts and several datasets. Dadi et al.[1] showed that across many cohorts and clinical questions, the tangent kind should be preferred.
plt.show()
References¶
Total running time of the script: (6 minutes 2.954 seconds)
Estimated memory usage: 1003 MB