Note
This page is a reference documentation. It only explains the function signature, and not how to use it. Please refer to the user guide for the big picture.
nilearn.datasets.fetch_atlas_difumo¶
- nilearn.datasets.fetch_atlas_difumo(dimension=64, resolution_mm=2, data_dir=None, resume=True, verbose=1)[source]¶
Fetch DiFuMo brain atlas.
Dictionaries of Functional Modes, or “DiFuMo”, can serve as probabilistic atlases to extract functional signals with different dimensionalities (64, 128, 256, 512, and 1024). These modes are optimized to represent well raw BOLD timeseries, over a with range of experimental conditions. See Dadi et al.[1].
Added in Nilearn 0.7.1.
- Parameters:
- dimension
int, default=64 Number of dimensions in the dictionary. Valid resolutions available are {64, 128, 256, 512, 1024}.
- resolution_mm
int, default=2mm The resolution in mm of the atlas to fetch. Valid options available are {2, 3}.
- data_dir
pathlib.Pathorstror None, optional Path where data should be downloaded. By default, files are downloaded in a
nilearn_datafolder in the home directory of the user. See alsonilearn.datasets.utils.get_data_dirs.- resume
bool, default=True Whether to resume download of a partly-downloaded file.
- verbose
boolorint, default=1 Verbosity level (
0orFalsemeans no message).
- dimension
- Returns:
- data
sklearn.utils.Bunch Dictionary-like object, the interest attributes are :
- ‘maps’:
str, path to 4D nifti file containing regions definition. The shape of the image is
(104, 123, 104, dimension)wheredimensionis the requested dimension of the atlas.
- ‘maps’:
- ‘labels’:
pandas.DataFramecontaining the labels of the regions. The length of the label array corresponds to the number of dimensions requested.
data.labels[i]is the label corresponding to volumeiin the ‘maps’ image.
- ‘labels’:
- ‘description’
str Description of the dataset.
- ‘description’
- ‘atlas_type’
str Type of atlas. See Probabilistic atlas and Deterministic atlas.
- ‘atlas_type’
- ‘template’
str The standardized space of analysis in which the atlas results are provided. When known it should be a valid template name taken from the spaces described in the BIDS specification.
- ‘template’
- data
Notes
If the dataset files are already present in the user’s Nilearn data directory, this fetcher will not re-download them. To force a fresh download, you can remove the existing dataset folder from your local Nilearn data directory.
For more details on how Nilearn stores datasets.
Direct download links from OSF:
References
Examples using nilearn.datasets.fetch_atlas_difumo¶
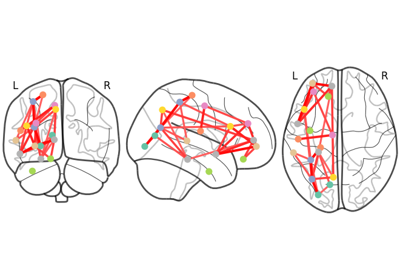
Comparing connectomes on different reference atlases