Note
This page is a reference documentation. It only explains the function signature, and not how to use it. Please refer to the user guide for the big picture.
nilearn.datasets.fetch_development_fmri¶
- nilearn.datasets.fetch_development_fmri(n_subjects=None, reduce_confounds=True, data_dir=None, resume=True, verbose=1, age_group='both')[source]¶
Fetch movie watching based brain development dataset (fMRI).
The data is downsampled to 4mm resolution for convenience with a repetition time (t_r) of 2 secs. The origin of the data is coming from OpenNeuro. See Notes below.
Please cite Richardson et al.[1] if you are using this dataset.
Added in Nilearn 0.5.2.
- Parameters:
- n_subjects
int, default=None The number of subjects to load. If None, all the subjects are loaded. Total 155 subjects.
- reduce_confounds
bool, default=True If True, the returned confounds only include 6 motion parameters, mean framewise displacement, signal from white matter, csf, and 6 anatomical compcor parameters. This selection only serves the purpose of having realistic examples. Depending on your research question, other confounds might be more appropriate. If False, returns all fMRIPrep confounds.
- data_dir
pathlib.Pathorstror None, optional Path where data should be downloaded. By default, files are downloaded in a
nilearn_datafolder in the home directory of the user. See alsonilearn.datasets.utils.get_data_dirs.- resume
bool, default=True Whether to resume download of a partly-downloaded file.
- verbose
boolorint, default=1 Verbosity level (
0orFalsemeans no message).- age_group
str, default=’both’ Which age group to fetch
‘adults’ = fetch adults only (n=33, ages 18-39)
‘child’ = fetch children only (n=122, ages 3-12)
‘both’ = fetch full sample (n=155)
- n_subjects
- Returns:
- dataBunch
Dictionary-like object, the interest attributes are :
- ‘phenotypic’: pandas.DataFame
Contains each subject age, age group, child or adult, gender, handedness.
- ‘t_r’:
float Repetition time of the functional data.
- ‘t_r’:
Notes
If the dataset files are already present in the user’s Nilearn data directory, this fetcher will not re-download them. To force a fresh download, you can remove the existing dataset folder from your local Nilearn data directory.
For more details on how Nilearn stores datasets.
The original data is downloaded from OpenNeuro https://openneuro.org/datasets/ds000228/versions/1.0.0
This fetcher downloads downsampled data that are available on Open Science Framework (OSF). Located here: https://osf.io/5hju4/files/
Preprocessing details: https://osf.io/wjtyq/
Note that if n_subjects > 2, and age_group is ‘both’, fetcher will return a ratio of children and adults representative of the total sample.
References
Examples using nilearn.datasets.fetch_development_fmri¶
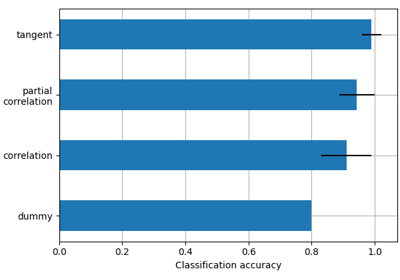
Classification of age groups using functional connectivity
Clustering methods to learn a brain parcellation from fMRI
Comparing connectomes on different reference atlases
Computing a connectome with sparse inverse covariance
Deriving spatial maps from group fMRI data using ICA and Dictionary Learning
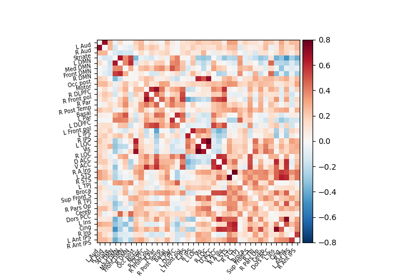
Extracting signals of a probabilistic atlas of functional regions
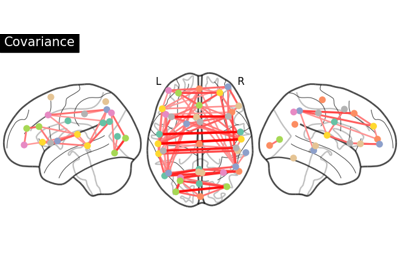
Group Sparse inverse covariance for multi-subject connectome
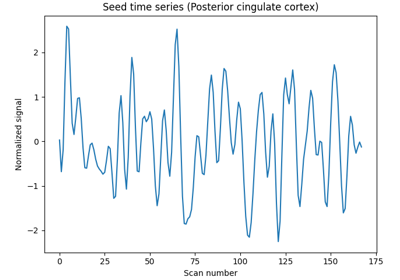
Producing single subject maps of seed-to-voxel correlation
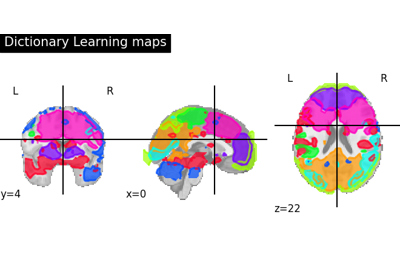
Regions extraction using dictionary learning and functional connectomes
Extracting signals from brain regions using the NiftiLabelsMasker
Multivariate decompositions: Independent component analysis of fMRI