Note
This page is a reference documentation. It only explains the function signature, and not how to use it. Please refer to the user guide for the big picture.
nilearn.datasets.fetch_megatrawls_netmats¶
- nilearn.datasets.fetch_megatrawls_netmats(dimensionality=100, timeseries='eigen_regression', matrices='partial_correlation', data_dir=None, resume=True, verbose=1)[source]¶
Download and return Network Matrices data from MegaTrawls release in HCP.
This data can be used to predict relationships between imaging data and non-imaging behavioral measures such as age, sex, education, etc. The network matrices are estimated from functional connectivity datasets of 461 subjects.
Technical details
For more technical details about predicting the measures, refer to: Stephen Smith et al, HCP beta-release of the Functional Connectivity MegaTrawl. April 2015 “HCP500-MegaTrawl” release. https://balsa.wustl.edu
Terms and conditions
This is open access data. You must agree to Terms and conditions of using this data before using it, available at: https://www.humanconnectome.org/study/hcp-young-adult/document/wu-minn-hcp-consortium-open-access-data-use-terms
- Parameters:
- dimensionality
int, default=100 Valid inputs are 25, 50, 100, 200, 300. By default, network matrices estimated using Group ICA brain parcellation of 100 components/dimensions will be returned.
- timeseries
str, default=’eigen_regression’ Valid inputs are ‘multiple_spatial_regression’ or ‘eigen_regression’. By default ‘eigen_regression’, matrices estimated using first principal eigen component timeseries signals extracted from each subject data parcellations will be returned. Otherwise, ‘multiple_spatial_regression’ matrices estimated using spatial regressor based timeseries signals extracted from each subject data parcellations will be returned.
- matrices
str, default=’partial_correlation’ Valid inputs are ‘full_correlation’ or ‘partial_correlation’. By default, partial correlation matrices will be returned otherwise if selected full correlation matrices will be returned.
- data_dir
pathlib.Pathorstror None, optional Path where data should be downloaded. By default, files are downloaded in a
nilearn_datafolder in the home directory of the user. See alsonilearn.datasets.utils.get_data_dirs.- resume
bool, default=True Whether to resume download of a partly-downloaded file.
- verbose
boolorint, default=1 Verbosity level (
0orFalsemeans no message).
- dimensionality
- Returns:
- dataBunch
Dictionary-like object, the attributes are :
‘dimensions’: int, consists of given input in dimensions.
‘timeseries’: str, consists of given input in timeseries method.
‘matrices’: str, consists of given type of specific matrices.
‘correlation_matrices’: pd.DataFrame consists of correlation matrices based on given type of matrices. Array size will depend on given dimensions (n, n).
‘description’: data description
Notes
If the dataset files are already present in the user’s Nilearn data directory, this fetcher will not re-download them. To force a fresh download, you can remove the existing dataset folder from your local Nilearn data directory.
For more details on how Nilearn stores datasets.
For more information see the dataset description.
Examples using nilearn.datasets.fetch_megatrawls_netmats¶
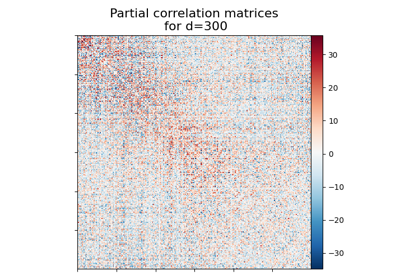
Visualizing Megatrawls Network Matrices from Human Connectome Project