Note
Go to the end to download the full example code. or to run this example in your browser via Binder
Extracting signals of a probabilistic atlas of functional regions¶
This example extracts the signal on regions defined via a probabilistic atlas, to construct a functional connectome.
We use the MSDL atlas of functional regions in movie-watching.
The key to extract signals is to use the
NiftiMapsMasker that can transform nifti
objects to time series using a probabilistic atlas.
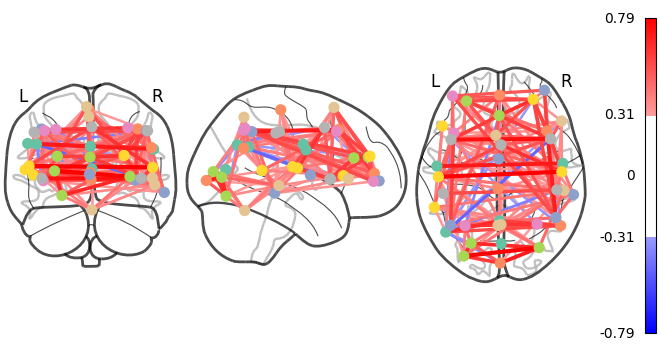
As the MSDL atlas comes with (x, y, z) MNI coordinates for the different regions, we can visualize the matrix as a graph of interaction in a brain. To avoid having too dense a graph, we represent only the 20% edges with the highest values.
Retrieve the atlas and the data¶
from nilearn.datasets import fetch_atlas_msdl, fetch_development_fmri
atlas = fetch_atlas_msdl()
# Loading atlas image stored in 'maps'
atlas_filename = atlas["maps"]
# Loading atlas data stored in 'labels'
labels = atlas["labels"]
# Load the functional datasets
data = fetch_development_fmri(n_subjects=1)
print(
"First subject resting-state nifti image (4D) is located "
f"at: {data.func[0]}"
)
[fetch_atlas_msdl] Dataset found in /home/runner/nilearn_data/msdl_atlas
[fetch_development_fmri] Dataset found in
/home/runner/nilearn_data/development_fmri
[fetch_development_fmri] Dataset found in
/home/runner/nilearn_data/development_fmri/development_fmri
[fetch_development_fmri] Dataset found in
/home/runner/nilearn_data/development_fmri/development_fmri
First subject resting-state nifti image (4D) is located at: /home/runner/nilearn_data/development_fmri/development_fmri/sub-pixar123_task-pixar_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz
Extract the time series¶
from nilearn.maskers import NiftiMapsMasker
masker = NiftiMapsMasker(
maps_img=atlas_filename,
standardize_confounds=True,
memory="nilearn_cache",
memory_level=1,
verbose=1,
)
time_series = masker.fit_transform(data.func[0], confounds=data.confounds)
\[NiftiMapsMasker.wrapped] Loading regions from
'/home/runner/nilearn_data/msdl_atlas/MSDL_rois/msdl_rois.nii'
\[NiftiMapsMasker.wrapped] Resampling regions
\[NiftiMapsMasker.wrapped] Finished fit
/home/runner/work/nilearn/nilearn/examples/03_connectivity/plot_probabilistic_atlas_extraction.py:56: FutureWarning:
boolean values for 'standardize' will be deprecated in nilearn 0.15.0.
Use 'zscore_sample' instead of 'True' or use 'None' instead of 'False'.
We can generate an HTML report and visualize the components of the
NiftiMapsMasker.
You can pass the indices of the spatial maps you want to include in the
report in the order you want them to appear.
Here, we only include maps 2, 6, 7, 16, and 21 in the report:
report = masker.generate_report(displayed_maps=[2, 6, 7, 16, 21])
report
time_series is now a 2D matrix, of shape (number of time points x number of regions)
print(time_series.shape)
(168, 39)
Build and display a correlation matrix¶
from nilearn.connectome import ConnectivityMeasure
correlation_measure = ConnectivityMeasure(kind="correlation", verbose=1)
correlation_matrix = correlation_measure.fit_transform([time_series])[0]
# Display the correlation matrix
import numpy as np
from nilearn.plotting import plot_connectome, plot_matrix, show
# Mask out the major diagonal
np.fill_diagonal(correlation_matrix, 0)
plot_matrix(correlation_matrix, labels=labels, vmax=0.8, vmin=-0.8)
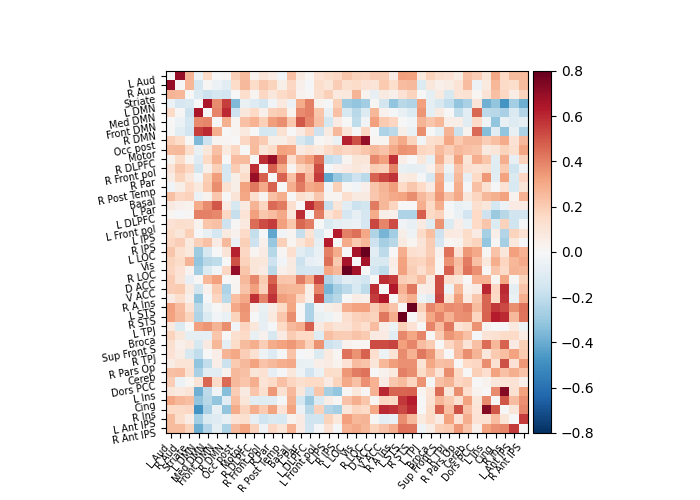
\[ConnectivityMeasure.wrapped] Finished fit
<matplotlib.image.AxesImage object at 0x7fe008d77e80>
And now display the corresponding graph¶
coords = atlas.region_coords
# We threshold to keep only the 20% of edges with the highest value
# because the graph is very dense
plot_connectome(correlation_matrix, coords, edge_threshold="80%")
show()
3D visualization in a web browser¶
An alternative to plot_connectome is to use
view_connectome that gives more interactive
visualizations in a web browser. See 3D Plots of connectomes
for more details.
from nilearn.plotting import view_connectome
view = view_connectome(correlation_matrix, coords, edge_threshold="80%")
# In a notebook, if ``view`` is the output of a cell, it will
# be displayed below the cell
view
# uncomment this to open the plot in a web browser:
# view.open_in_browser()
Total running time of the script: (0 minutes 6.371 seconds)
Estimated memory usage: 130 MB