Note
Go to the end to download the full example code. or to run this example in your browser via Binder
Setting a parameter by cross-validation¶
Here we set the number of features selected in an Anova-SVC approach to maximize the cross-validation score.
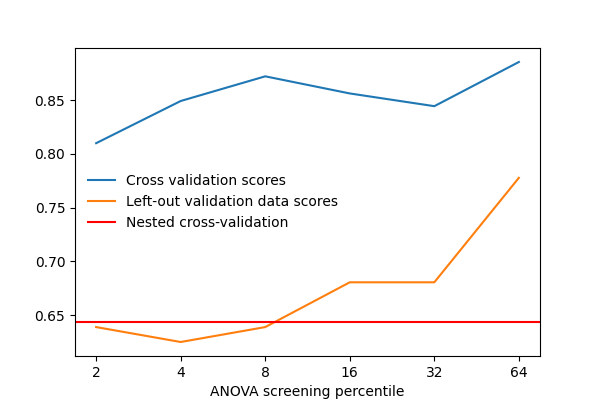
After separating 2 runs for validation, we vary that parameter and measure the cross-validation score. We also measure the prediction score on the left-out validation data. As we can see, the two scores vary by a significant amount: this is due to sampling noise in cross validation, and choosing the parameter k to maximize the cross-validation score, might not maximize the score on left-out data.
Thus using data to maximize a cross-validation score computed on that same data is likely to be too optimistic and lead to an overfit.
The proper approach is known as a “nested cross-validation”. It consists in doing cross-validation loops to set the model parameters inside the cross-validation loop used to judge the prediction performance: the parameters are set separately on each fold, never using the data used to measure performance.
For decoding tasks, in nilearn, this can be done using the
Decoder object, which will automatically select
the best parameters of an estimator from a grid of parameter values.
One difficulty is that the Decoder object is a composite estimator: a pipeline of feature selection followed by Support Vector Machine. Tuning the SVM’s parameters is already done automatically inside the Decoder, but performing cross-validation for the feature selection must be done manually.
import warnings
warnings.filterwarnings(
"ignore", message="The provided image has no sform in its header."
)
# set overall verbosity for this example
verbose = 2
Load the Haxby dataset¶
from nilearn import datasets
from nilearn.plotting import show
# by default 2nd subject data will be fetched on which we run our analysis
haxby_dataset = datasets.fetch_haxby()
fmri_img = haxby_dataset.func[0]
mask_img = haxby_dataset.mask
# print basic information on the dataset
print(f"Mask nifti image (3D) is located at: {haxby_dataset.mask}")
print(f"Functional nifti image (4D) are located at: {haxby_dataset.func[0]}")
# Load the behavioral data
import pandas as pd
labels = pd.read_csv(haxby_dataset.session_target[0], sep=" ")
y = labels["labels"]
# Keep only data corresponding to shoes or bottles
from nilearn.image import index_img
condition_mask = y.isin(["shoe", "bottle"])
fmri_niimgs = index_img(fmri_img, condition_mask)
y = y[condition_mask]
run = labels["chunks"][condition_mask]
[fetch_haxby] Dataset found in /home/runner/nilearn_data/haxby2001
Mask nifti image (3D) is located at: /home/runner/nilearn_data/haxby2001/mask.nii.gz
Functional nifti image (4D) are located at: /home/runner/nilearn_data/haxby2001/subj2/bold.nii.gz
ANOVA pipeline with Decoder object¶
Nilearn Decoder object aims to provide smooth user experience by acting as a pipeline of several tasks: preprocessing with NiftiMasker, reducing dimension by selecting only relevant features with ANOVA – a classical univariate feature selection based on F-test, and then decoding with different types of estimators (in this example is Support Vector Machine with a linear kernel) on nested cross-validation.
from nilearn.decoding import Decoder
# We provide a grid of hyperparameter values to the Decoder's internal
# cross-validation. If no param_grid is provided, the Decoder will use a
# default grid with sensible values for the chosen estimator
param_grid = [
{
"penalty": ["l2"],
"dual": [True],
"C": [100, 1000],
},
{
"penalty": ["l1"],
"dual": [False],
"C": [100, 1000],
},
]
# Here screening_percentile is set to 2 percent, meaning around 800
# features will be selected with ANOVA.
decoder = Decoder(
estimator="svc",
cv=5,
mask=mask_img,
smoothing_fwhm=4,
screening_percentile=2,
param_grid=param_grid,
verbose=verbose,
)
Fit the Decoder and predict the responses¶
As a complete pipeline by itself, decoder will perform cross-validation for the estimator, in this case Support Vector Machine. We can output the best parameters selected for each cross-validation fold. See https://scikit-learn.org/stable/modules/cross_validation.html for an excellent explanation of how cross-validation works.
# Fit the Decoder
decoder.fit(fmri_niimgs, y)
# Print the best parameters for each fold
for i, (best_c, best_penalty, best_dual, cv_score) in enumerate(
zip(
decoder.cv_params_["shoe"]["C"],
decoder.cv_params_["shoe"]["penalty"],
decoder.cv_params_["shoe"]["dual"],
decoder.cv_scores_["shoe"],
strict=False,
)
):
print(
f"Fold {i + 1} | Best SVM parameters: C={best_c}"
f", penalty={best_penalty}, dual={best_dual} with score: {cv_score}"
)
# Output the prediction with Decoder
y_pred = decoder.predict(fmri_niimgs)
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023abec50>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:126: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023abec50>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 2
\[Decoder.fit] Corrected screening-percentile: 1.9171
\[Decoder.fit] The decoding model will be trained on 399 features.
\[Decoder.fit] The decoding model will be trained on 399 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 400 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.7s remaining: 0.0s
\[Decoder.fit] Selection kept 400 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 1.4s remaining: 0.0s
\[Decoder.fit] Selection kept 400 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 2.1s remaining: 0.0s
\[Decoder.fit] Selection kept 400 features.
\[Decoder.fit] Selection kept 400 features.
[Parallel(n_jobs=1)]: Done 5 out of 5 | elapsed: 3.5s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
Fold 1 | Best SVM parameters: C=1000, penalty=l1, dual=False with score: 0.9483471074380164
Fold 2 | Best SVM parameters: C=1000, penalty=l2, dual=True with score: 0.9177489177489176
Fold 3 | Best SVM parameters: C=100, penalty=l1, dual=False with score: 0.7489177489177489
Fold 4 | Best SVM parameters: C=100, penalty=l1, dual=False with score: 0.7792207792207793
Fold 5 | Best SVM parameters: C=1000, penalty=l1, dual=False with score: 0.7597402597402597
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023abec50>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
Compute prediction scores with different values of screening percentile¶
import numpy as np
screening_percentile_range = [2, 4, 8, 16, 32, 64]
cv_scores = []
val_scores = []
for sp in screening_percentile_range:
print("\n")
decoder = Decoder(
estimator="svc",
mask=mask_img,
smoothing_fwhm=4,
cv=3,
screening_percentile=sp,
param_grid=param_grid,
verbose=verbose,
)
decoder.fit(index_img(fmri_niimgs, run < 10), y[run < 10])
cv_scores.append(np.mean(decoder.cv_scores_["bottle"]))
print(f"Sreening Percentile: {sp:.3f}")
print(f"Mean CV score: {cv_scores[-1]:.4f}")
y_pred = decoder.predict(index_img(fmri_niimgs, run == 10))
val_scores.append(np.mean(y_pred == y[run == 10]))
print(f"Validation score: {val_scores[-1]:.4f}")
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe002d8f370>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:166: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe002d8f370>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 2
\[Decoder.fit] Corrected screening-percentile: 1.9171
\[Decoder.fit] The decoding model will be trained on 399 features.
\[Decoder.fit] The decoding model will be trained on 399 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 400 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.4s remaining: 0.0s
\[Decoder.fit] Selection kept 400 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 0.9s remaining: 0.0s
\[Decoder.fit] Selection kept 400 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 1.4s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 1.4s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
Sreening Percentile: 2.000
Mean CV score: 0.8300
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe002d8ed70>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
Validation score: 0.4444
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe002d8db70>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:166: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe002d8db70>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 4
\[Decoder.fit] Corrected screening-percentile: 3.83419
\[Decoder.fit] The decoding model will be trained on 1197 features.
\[Decoder.fit] The decoding model will be trained on 1197 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 1198 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.7s remaining: 0.0s
\[Decoder.fit] Selection kept 1198 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 1.4s remaining: 0.0s
\[Decoder.fit] Selection kept 1198 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 2.1s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 2.1s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
Sreening Percentile: 4.000
Mean CV score: 0.8493
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe002d8cca0>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
Validation score: 0.3889
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fdfdf7acf40>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:166: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fdfdf7acf40>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 8
\[Decoder.fit] Corrected screening-percentile: 7.66838
\[Decoder.fit] The decoding model will be trained on 2793 features.
\[Decoder.fit] The decoding model will be trained on 2793 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 2794 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 1.0s remaining: 0.0s
\[Decoder.fit] Selection kept 2794 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 2.0s remaining: 0.0s
\[Decoder.fit] Selection kept 2794 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 2.8s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 2.8s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
Sreening Percentile: 8.000
Mean CV score: 0.8700
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fdfdf7acfd0>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
Validation score: 0.3889
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fdfdf7afd00>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:166: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fdfdf7afd00>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 16
\[Decoder.fit] Corrected screening-percentile: 15.3368
\[Decoder.fit] The decoding model will be trained on 5986 features.
\[Decoder.fit] The decoding model will be trained on 5986 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 5987 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 1.2s remaining: 0.0s
\[Decoder.fit] Selection kept 5987 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 2.7s remaining: 0.0s
\[Decoder.fit] Selection kept 5987 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 4.0s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 4.0s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
Sreening Percentile: 16.000
Mean CV score: 0.8637
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe002d8c100>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
Validation score: 0.7222
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe002d8db70>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:166: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe002d8db70>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 32
\[Decoder.fit] Corrected screening-percentile: 30.6735
\[Decoder.fit] The decoding model will be trained on 11973 features.
\[Decoder.fit] The decoding model will be trained on 11973 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 11974 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 2.0s remaining: 0.0s
\[Decoder.fit] Selection kept 11974 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 4.3s remaining: 0.0s
\[Decoder.fit] Selection kept 11974 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 5.9s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 5.9s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
Sreening Percentile: 32.000
Mean CV score: 0.8733
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fdfe0ea9b70>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
Validation score: 0.4444
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023abfc70>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:166: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023abfc70>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 64
\[Decoder.fit] Corrected screening-percentile: 61.3471
\[Decoder.fit] The decoding model will be trained on 24346 features.
\[Decoder.fit] The decoding model will be trained on 24346 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 24346 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 3.6s remaining: 0.0s
\[Decoder.fit] Selection kept 24346 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 7.1s remaining: 0.0s
\[Decoder.fit] Selection kept 24346 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 10.2s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 10.2s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
Sreening Percentile: 64.000
Mean CV score: 0.8722
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023abe470>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
Validation score: 0.3333
Nested cross-validation¶
We are going to tune the parameter ‘screening_percentile’ in the pipeline.
Note
We increase the tolerance a bit to make it easier for the fitting to converge.
from sklearn.model_selection import KFold
cv = KFold(n_splits=3)
nested_cv_scores = []
for train, test in cv.split(run):
y_train = np.array(y)[train]
y_test = np.array(y)[test]
val_scores = []
for sp in screening_percentile_range:
decoder = Decoder(
estimator="svc",
mask=mask_img,
smoothing_fwhm=4,
cv=3,
standardize="zscore_sample",
screening_percentile=sp,
param_grid=param_grid,
verbose=verbose,
estimator_args={"tol": 0.0005},
)
decoder.fit(index_img(fmri_niimgs, train), y_train)
y_pred = decoder.predict(index_img(fmri_niimgs, test))
val_scores.append(np.mean(y_pred == y_test))
nested_cv_scores.append(np.max(val_scores))
print(f"Nested CV score: {np.mean(nested_cv_scores):.4f}")
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fdfdf7aeb00>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:209: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fdfdf7aeb00>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 2
\[Decoder.fit] Corrected screening-percentile: 1.9171
\[Decoder.fit] The decoding model will be trained on 399 features.
\[Decoder.fit] The decoding model will be trained on 399 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 400 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.2s remaining: 0.0s
\[Decoder.fit] Selection kept 400 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 0.3s remaining: 0.0s
\[Decoder.fit] Selection kept 400 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 0.4s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 0.4s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fdfdf7ac220>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023abf3d0>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:209: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023abf3d0>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 4
\[Decoder.fit] Corrected screening-percentile: 3.83419
\[Decoder.fit] The decoding model will be trained on 1197 features.
\[Decoder.fit] The decoding model will be trained on 1197 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 1198 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.2s remaining: 0.0s
\[Decoder.fit] Selection kept 1198 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 0.4s remaining: 0.0s
\[Decoder.fit] Selection kept 1198 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 0.6s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 0.6s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023abcac0>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023abe5f0>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:209: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023abe5f0>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 8
\[Decoder.fit] Corrected screening-percentile: 7.66838
\[Decoder.fit] The decoding model will be trained on 2793 features.
\[Decoder.fit] The decoding model will be trained on 2793 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 2794 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.3s remaining: 0.0s
\[Decoder.fit] Selection kept 2794 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 0.6s remaining: 0.0s
\[Decoder.fit] Selection kept 2794 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 0.8s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 0.8s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fdfdf7af490>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fdfdf7ac850>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:209: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fdfdf7ac850>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 16
\[Decoder.fit] Corrected screening-percentile: 15.3368
\[Decoder.fit] The decoding model will be trained on 5986 features.
\[Decoder.fit] The decoding model will be trained on 5986 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 5987 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.5s remaining: 0.0s
\[Decoder.fit] Selection kept 5987 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 0.9s remaining: 0.0s
\[Decoder.fit] Selection kept 5987 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 1.3s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 1.3s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023abfa30>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023abc3d0>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:209: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023abc3d0>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 32
\[Decoder.fit] Corrected screening-percentile: 30.6735
\[Decoder.fit] The decoding model will be trained on 11973 features.
\[Decoder.fit] The decoding model will be trained on 11973 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 11974 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.8s remaining: 0.0s
\[Decoder.fit] Selection kept 11974 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 1.4s remaining: 0.0s
\[Decoder.fit] Selection kept 11974 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 2.0s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 2.0s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023abc3d0>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe002d8db70>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:209: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe002d8db70>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 64
\[Decoder.fit] Corrected screening-percentile: 61.3471
\[Decoder.fit] The decoding model will be trained on 24346 features.
\[Decoder.fit] The decoding model will be trained on 24346 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 24346 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 1.5s remaining: 0.0s
\[Decoder.fit] Selection kept 24346 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 2.9s remaining: 0.0s
\[Decoder.fit] Selection kept 24346 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 4.1s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 4.1s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe002d8e680>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe008d77550>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:209: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe008d77550>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 2
\[Decoder.fit] Corrected screening-percentile: 1.9171
\[Decoder.fit] The decoding model will be trained on 399 features.
\[Decoder.fit] The decoding model will be trained on 399 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 400 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.1s remaining: 0.0s
\[Decoder.fit] Selection kept 400 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 0.2s remaining: 0.0s
\[Decoder.fit] Selection kept 400 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 0.4s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 0.4s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe008d75c30>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe008d76fb0>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:209: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe008d76fb0>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 4
\[Decoder.fit] Corrected screening-percentile: 3.83419
\[Decoder.fit] The decoding model will be trained on 1197 features.
\[Decoder.fit] The decoding model will be trained on 1197 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 1198 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.2s remaining: 0.0s
\[Decoder.fit] Selection kept 1198 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 0.4s remaining: 0.0s
\[Decoder.fit] Selection kept 1198 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 0.6s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 0.6s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe008d770a0>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe008d75210>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:209: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe008d75210>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 8
\[Decoder.fit] Corrected screening-percentile: 7.66838
\[Decoder.fit] The decoding model will be trained on 2793 features.
\[Decoder.fit] The decoding model will be trained on 2793 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 2794 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.3s remaining: 0.0s
\[Decoder.fit] Selection kept 2794 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 0.7s remaining: 0.0s
\[Decoder.fit] Selection kept 2794 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 1.0s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 1.0s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe002d8cf40>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe002d8ed70>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:209: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe002d8ed70>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 16
\[Decoder.fit] Corrected screening-percentile: 15.3368
\[Decoder.fit] The decoding model will be trained on 5986 features.
\[Decoder.fit] The decoding model will be trained on 5986 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 5987 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.5s remaining: 0.0s
\[Decoder.fit] Selection kept 5987 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 0.9s remaining: 0.0s
\[Decoder.fit] Selection kept 5987 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 1.5s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 1.5s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023abc970>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fdfdf7af4f0>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:209: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fdfdf7af4f0>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 32
\[Decoder.fit] Corrected screening-percentile: 30.6735
\[Decoder.fit] The decoding model will be trained on 11973 features.
\[Decoder.fit] The decoding model will be trained on 11973 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 11974 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.7s remaining: 0.0s
\[Decoder.fit] Selection kept 11974 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 1.5s remaining: 0.0s
\[Decoder.fit] Selection kept 11974 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 2.4s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 2.4s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023a11090>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023a10910>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:209: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023a10910>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 64
\[Decoder.fit] Corrected screening-percentile: 61.3471
\[Decoder.fit] The decoding model will be trained on 24346 features.
\[Decoder.fit] The decoding model will be trained on 24346 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 24346 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 1.6s remaining: 0.0s
\[Decoder.fit] Selection kept 24346 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 3.0s remaining: 0.0s
\[Decoder.fit] Selection kept 24346 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 4.7s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 4.7s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023a13970>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023a102b0>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:209: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023a102b0>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 2
\[Decoder.fit] Corrected screening-percentile: 1.9171
\[Decoder.fit] The decoding model will be trained on 399 features.
\[Decoder.fit] The decoding model will be trained on 399 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 400 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.1s remaining: 0.0s
\[Decoder.fit] Selection kept 400 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 0.3s remaining: 0.0s
\[Decoder.fit] Selection kept 400 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 0.4s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 0.4s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe030593190>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023a11bd0>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:209: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023a11bd0>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 4
\[Decoder.fit] Corrected screening-percentile: 3.83419
\[Decoder.fit] The decoding model will be trained on 1197 features.
\[Decoder.fit] The decoding model will be trained on 1197 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 1198 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.2s remaining: 0.0s
\[Decoder.fit] Selection kept 1198 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 0.4s remaining: 0.0s
\[Decoder.fit] Selection kept 1198 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 0.6s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 0.6s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023a11e70>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023a125f0>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:209: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023a125f0>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 8
\[Decoder.fit] Corrected screening-percentile: 7.66838
\[Decoder.fit] The decoding model will be trained on 2793 features.
\[Decoder.fit] The decoding model will be trained on 2793 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 2794 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.2s remaining: 0.0s
\[Decoder.fit] Selection kept 2794 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 0.6s remaining: 0.0s
\[Decoder.fit] Selection kept 2794 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 0.9s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 0.9s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023abc880>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023a11f00>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:209: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe023a11f00>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 16
\[Decoder.fit] Corrected screening-percentile: 15.3368
\[Decoder.fit] The decoding model will be trained on 5986 features.
\[Decoder.fit] The decoding model will be trained on 5986 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 5987 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.4s remaining: 0.0s
\[Decoder.fit] Selection kept 5987 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 0.9s remaining: 0.0s
\[Decoder.fit] Selection kept 5987 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 1.3s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 1.3s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe008d77010>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe008d767a0>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:209: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fe008d767a0>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 32
\[Decoder.fit] Corrected screening-percentile: 30.6735
\[Decoder.fit] The decoding model will be trained on 11973 features.
\[Decoder.fit] The decoding model will be trained on 11973 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 11974 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.7s remaining: 0.0s
\[Decoder.fit] Selection kept 11974 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 1.4s remaining: 0.0s
\[Decoder.fit] Selection kept 11974 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 2.0s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 2.0s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fdfe2051930>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
\[Decoder.fit] Loading mask from
'/home/runner/nilearn_data/haxby2001/mask.nii.gz'
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fdfe2052dd0>
/home/runner/work/nilearn/nilearn/examples/02_decoding/plot_haxby_grid_search.py:209: UserWarning:
[NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
\[Decoder.fit] Resampling mask
\[Decoder.fit] Finished fit
\[Decoder.fit] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fdfe2052dd0>
\[Decoder.fit] Smoothing images
\[Decoder.fit] Extracting region signals
\[Decoder.fit] Cleaning extracted signals
\[Decoder.fit] Mask volume = 1.96442e+06mm^3 = 1964.42cm^3
\[Decoder.fit] Standard brain volume = 1.88299e+06mm^3
\[Decoder.fit] Original screening-percentile: 64
\[Decoder.fit] Corrected screening-percentile: 61.3471
\[Decoder.fit] The decoding model will be trained on 24346 features.
\[Decoder.fit] The decoding model will be trained on 24346 features.
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
\[Decoder.fit] Selection kept 24346 features.
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 1.6s remaining: 0.0s
\[Decoder.fit] Selection kept 24346 features.
[Parallel(n_jobs=1)]: Done 2 out of 2 | elapsed: 3.1s remaining: 0.0s
\[Decoder.fit] Selection kept 24346 features.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 4.7s remaining: 0.0s
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 4.7s finished
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.fit] Computing image from signals
\[Decoder.predict] Loading data from <nibabel.nifti1.Nifti1Image object at
0x7fdfe2053d00>
\[Decoder.predict] Smoothing images
\[Decoder.predict] Extracting region signals
\[Decoder.predict] Cleaning extracted signals
Nested CV score: 0.6528
Plot the prediction scores using matplotlib¶
from matplotlib import pyplot as plt
plt.figure(figsize=(6, 4))
plt.plot(cv_scores, label="Cross validation scores")
plt.plot(val_scores, label="Left-out validation data scores")
plt.xticks(
np.arange(len(screening_percentile_range)), screening_percentile_range
)
plt.axis("tight")
plt.xlabel("ANOVA screening percentile")
plt.axhline(
np.mean(nested_cv_scores), label="Nested cross-validation", color="r"
)
plt.legend(loc="best", frameon=False)
show()
Total running time of the script: (1 minutes 37.081 seconds)
Estimated memory usage: 1210 MB