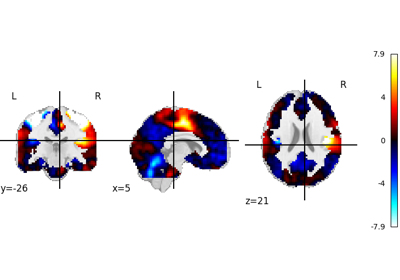
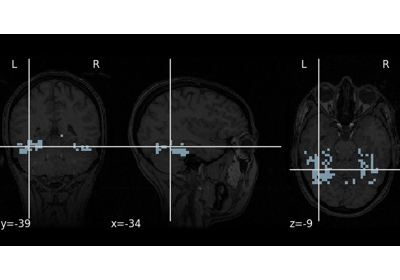
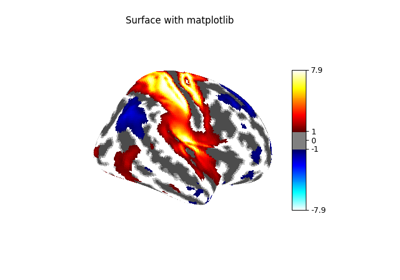
1. Introduction¶
2. What is nilearn?¶
nilearn is a package that makes it easy to use advanced machine learning techniques
to analyze data acquired with MRI machines.
In particular, underlying machine learning problems include
decoding brain data,
computing brain parcellations,
analyzing functional connectivity and connectomes,
doing multi-voxel pattern analysis (MVPA) or predictive modeling.
nilearn can readily be used on task fMRI,
resting-state, or
voxel-based morphometry (VBM) data.
For machine learning experts, the value of nilearn can be seen as
domain-specific feature engineering construction, that is, shaping
neuroimaging data into a feature matrix well suited for statistical learning.
Note
It is ok if these terms don’t make sense to you yet: this guide will walk you through them in a comprehensive manner.
3. Using nilearn for the first time¶
nilearn is a Python library. If you have never used Python before,
you should probably have a look at a general introduction about Python
as well as to Scientific Python Lectures before diving into nilearn.
3.1. First steps with nilearn¶
At this stage, you should have installed nilearn and opened a Jupyter notebook
or an IPython / Python session. First, load nilearn with
import nilearn
nilearn comes in with some data that are commonly used in neuroimaging.
For instance, it comes with volumic template images of brains such as MNI:
print(nilearn.datasets.MNI152_FILE_PATH)
Output:
'/home/yasmin/nilearn/nilearn/nilearn/datasets/data/mni_icbm152_t1_tal_nlin_sym_09a_converted.nii.gz'
Let’s have a look at this image:
nilearn.plotting.plot_img(nilearn.datasets.MNI152_FILE_PATH)
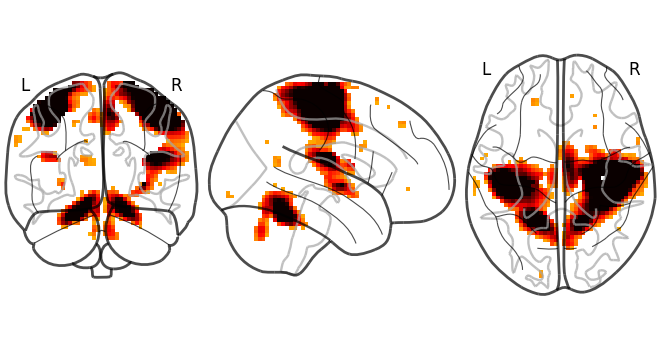
3.2. Learning with the API references¶
In the last command, you just made use of 2 nilearn modules: nilearn.datasets
and nilearn.plotting.
All modules are described in the API references.
Oftentimes, if you are already familiar with the problems and vocabulary of MRI analysis,
the module and function names are explicit enough that you should understand what nilearn does.
Note
Exercise: Varying the amount of smoothing in an image
Compute the mean EPI for one individual of the brain development
dataset downloaded with nilearn.datasets.fetch_development_fmri and
smooth it with an FWHM varying from 0mm to 20mm in increments of 5mm
Intermediate steps:
Run
nilearn.datasets.fetch_development_fmriand inspect the.keys()of the returned objectCheck the
nilearn.imagemodule in the documentation to find a function to compute the mean of a 4D imageCheck the
nilearn.imagemodule again to find a function which smoothes imagesPlot the computed image for each smoothing value
A solution can be found here.
3.3. Learning with examples¶
nilearn comes with a lot of examples/tutorials.
Going through them should give you a precise overview of what you can achieve with this package.
For new-comers, we recommend going through the following examples in the suggested order:
Basic nilearn example: manipulating and looking at data
Intro to GLM Analysis: a single-run, single-subject fMRI dataset
Computing a Region of Interest (ROI) mask manually
Classification of age groups using functional connectivity
3.4. Finding help¶
On top of this guide, there is a lot of content available outside of nilearn
that could be of interest to new-comers:
Handbook of Functional MRI Data Analysis by Russel Poldrack, Jeanette Mumford and Thomas Nichols.
The documentation of
scikit-learnexplains each method with tips on practical use and examples: https://scikit-learn.org/stable/. While not specific to neuroimaging, it is often a recommended read.(For Python beginners) A quick and gentle introduction to scientific computing with Python with the scientififc Python lectures. Moreover, you can use
nilearnwith Jupyter notebooks or IPython sessions. They provide an interactive environment that greatly facilitates debugging and visualization.
Besides, you can find help on neurostars for questions
related to nilearn and to computational neuroscience in general.
Finally, the nilearn team organizes weekly drop-in hours.
We can also be reached on github
in case you find a bug.
4. Machine learning applications to Neuroimaging¶
nilearn brings easy-to-use machine learning tools that can be leveraged to solve more complex applications.
The interested reader can dive into the following articles for more content.
We give a non-exhaustive list of such important applications.
Diagnosis and prognosis
Predicting a clinical score or even treatment response from brain imaging with supervised learning e.g. Mourão-Miranda et al.[1].
Information mapping
Using the prediction accuracy of a classifier to characterize relationships between brain images and stimuli. (e.g. searchlight and Kriegeskorte et al.[2])
Transfer learning
Measuring how much an estimator trained on one specific psychological process/task can predict the neural activity underlying another specific psychological process/task (e.g. discriminating left from right eye movements also discriminates additions from subtractions [3])
High-dimensional multivariate statistics
From a statistical point of view, machine learning implements statistical estimation of models with a large number of parameters. Tricks pulled in machine learning (e.g. regularization) can make this estimation possible despite the usually small number of observations in the neuroimaging domain [4]. This usage of machine learning requires some understanding of the models.
Data mining / exploration
Data-driven exploration of brain images. This includes the extraction of the major brain networks from resting-state data (“resting-state networks”) or movie-watching data as well as the discovery of connectionally coherent functional modules (“connectivity-based parcellation”). For example, Extracting functional brain networks: ICA and related or Clustering to parcellate the brain in regions with clustering.