Note
Click here to download the full example code or to run this example in your browser via Binder
9.2.16. Seed-based connectivity on the surface¶
The dataset that is a subset of the enhanced NKI Rockland sample (http://fcon_1000.projects.nitrc.org/indi/enhanced/, Nooner et al, 2012)
Resting state fMRI scans (TR=645ms) of 102 subjects were preprocessed (https://github.com/fliem/nki_nilearn) and projected onto the Freesurfer fsaverage5 template (Dale et al, 1999, Fischl et al, 1999). For this example we use the time series of a single subject’s left hemisphere.
The Destrieux parcellation (Destrieux et al, 2010) in fsaverage5 space as distributed with Freesurfer is used to select a seed region in the posterior cingulate cortex.
Functional connectivity of the seed region to all other cortical nodes in the same hemisphere is calculated using Pearson product-moment correlation coefficient.
The nilearn.plotting.plot_surf_stat_map function is used to plot the resulting statistical map on the (inflated) pial surface.
See also for a similar example but using volumetric input data.
See Plotting brain images for more details on plotting tools.
9.2.16.1. References¶
Nooner et al, (2012). The NKI-Rockland Sample: A model for accelerating the pace of discovery science in psychiatry. Frontiers in neuroscience 6, 152. URL http://dx.doi.org/10.3389/fnins.2012.00152
Dale et al, (1999). Cortical surface-based analysis.I. Segmentation and surface reconstruction. Neuroimage 9. URL http://dx.doi.org/10.1006/nimg.1998.0395
Fischl et al, (1999). Cortical surface-based analysis. II: Inflation, flattening, and a surface-based coordinate system. Neuroimage 9. http://dx.doi.org/10.1006/nimg.1998.0396
Destrieux et al, (2010). Automatic parcellation of human cortical gyri and sulci using standard anatomical nomenclature. NeuroImage, 53, 1. URL http://dx.doi.org/10.1016/j.neuroimage.2010.06.010.
9.2.16.2. Retrieving the data¶
# NKI resting state data from nilearn
from nilearn import datasets
nki_dataset = datasets.fetch_surf_nki_enhanced(n_subjects=1)
# The nki dictionary contains file names for the data
# of all downloaded subjects.
print(('Resting state data of the first subjects on the '
'fsaverag5 surface left hemisphere is at: %s' %
nki_dataset['func_left'][0]))
# Destrieux parcellation for left hemisphere in fsaverage5 space
destrieux_atlas = datasets.fetch_atlas_surf_destrieux()
parcellation = destrieux_atlas['map_left']
labels = destrieux_atlas['labels']
# Fsaverage5 surface template
fsaverage = datasets.fetch_surf_fsaverage()
# The fsaverage dataset contains file names pointing to
# the file locations
print('Fsaverage5 pial surface of left hemisphere is at: %s' %
fsaverage['pial_left'])
print('Fsaverage5 inflated surface of left hemisphere is at: %s' %
fsaverage['infl_left'])
print('Fsaverage5 sulcal depth map of left hemisphere is at: %s' %
fsaverage['sulc_left'])
Out:
Dataset created in /home/varoquau/nilearn_data/nki_enhanced_surface
Downloading data from https://www.nitrc.org/frs/download.php/8470/pheno_nki_nilearn.csv ...
...done. (1 seconds, 0 min)
Downloading data from https://www.nitrc.org/frs/download.php/8261/A00028185_rh_preprocessed_fsaverage5_fwhm6.gii ...
Downloaded 106496 of 42412120 bytes (0.3%, 8.1min remaining)
Downloaded 286720 of 42412120 bytes (0.7%, 6.0min remaining)
Downloaded 581632 of 42412120 bytes (1.4%, 4.4min remaining)
Downloaded 999424 of 42412120 bytes (2.4%, 3.4min remaining)
Downloaded 1597440 of 42412120 bytes (3.8%, 2.6min remaining)
Downloaded 2326528 of 42412120 bytes (5.5%, 2.1min remaining)
Downloaded 3203072 of 42412120 bytes (7.6%, 1.7min remaining)
Downloaded 4276224 of 42412120 bytes (10.1%, 1.4min remaining)
Downloaded 6135808 of 42412120 bytes (14.5%, 1.0min remaining)
Downloaded 8650752 of 42412120 bytes (20.4%, 45.0s remaining)
Downloaded 12124160 of 42412120 bytes (28.6%, 31.3s remaining)
Downloaded 17072128 of 42412120 bytes (40.3%, 20.1s remaining)
Downloaded 23945216 of 42412120 bytes (56.5%, 11.2s remaining)
Downloaded 33439744 of 42412120 bytes (78.8%, 4.2s remaining) ...done. (18 seconds, 0 min)
Downloading data from https://www.nitrc.org/frs/download.php/8260/A00028185_lh_preprocessed_fsaverage5_fwhm6.gii ...
Downloaded 114688 of 42402060 bytes (0.3%, 7.5min remaining)
Downloaded 335872 of 42402060 bytes (0.8%, 5.1min remaining)
Downloaded 696320 of 42402060 bytes (1.6%, 3.7min remaining)
Downloaded 1146880 of 42402060 bytes (2.7%, 2.9min remaining)
Downloaded 1703936 of 42402060 bytes (4.0%, 2.4min remaining)
Downloaded 2400256 of 42402060 bytes (5.7%, 2.0min remaining)
Downloaded 3244032 of 42402060 bytes (7.7%, 1.7min remaining)
Downloaded 4292608 of 42402060 bytes (10.1%, 1.5min remaining)
Downloaded 5537792 of 42402060 bytes (13.1%, 1.2min remaining)
Downloaded 6676480 of 42402060 bytes (15.7%, 1.1min remaining)
Downloaded 8691712 of 42402060 bytes (20.5%, 51.4s remaining)
Downloaded 10256384 of 42402060 bytes (24.2%, 45.4s remaining)
Downloaded 11714560 of 42402060 bytes (27.6%, 41.2s remaining)
Downloaded 13287424 of 42402060 bytes (31.3%, 37.1s remaining)
Downloaded 14974976 of 42402060 bytes (35.3%, 33.2s remaining)
Downloaded 16793600 of 42402060 bytes (39.6%, 29.6s remaining)
Downloaded 18513920 of 42402060 bytes (43.7%, 26.6s remaining)
Downloaded 19972096 of 42402060 bytes (47.1%, 24.6s remaining)
Downloaded 21536768 of 42402060 bytes (50.8%, 22.4s remaining)
Downloaded 23142400 of 42402060 bytes (54.6%, 20.2s remaining)
Downloaded 24748032 of 42402060 bytes (58.4%, 18.2s remaining)
Downloaded 26378240 of 42402060 bytes (62.2%, 16.3s remaining)
Downloaded 28049408 of 42402060 bytes (66.2%, 14.3s remaining)
Downloaded 29827072 of 42402060 bytes (70.3%, 12.3s remaining)
Downloaded 31457280 of 42402060 bytes (74.2%, 10.6s remaining)
Downloaded 32849920 of 42402060 bytes (77.5%, 9.2s remaining)
Downloaded 34349056 of 42402060 bytes (81.0%, 7.7s remaining)
Downloaded 35962880 of 42402060 bytes (84.8%, 6.1s remaining)
Downloaded 37715968 of 42402060 bytes (88.9%, 4.4s remaining)
Downloaded 39460864 of 42402060 bytes (93.1%, 2.7s remaining)
Downloaded 40386560 of 42402060 bytes (95.2%, 1.9s remaining)
Downloaded 41615360 of 42402060 bytes (98.1%, 0.7s remaining) ...done. (41 seconds, 0 min)
Resting state data of the first subjects on the fsaverag5 surface left hemisphere is at: /home/varoquau/nilearn_data/nki_enhanced_surface/A00028185/A00028185_left_preprocessed_fwhm6.gii
Fsaverage5 pial surface of left hemisphere is at: /home/varoquau/dev/nilearn/nilearn/datasets/data/fsaverage5/pial.left.gii.gz
Fsaverage5 inflated surface of left hemisphere is at: /home/varoquau/dev/nilearn/nilearn/datasets/data/fsaverage5/pial_inflated.left.gii.gz
Fsaverage5 sulcal depth map of left hemisphere is at: /home/varoquau/dev/nilearn/nilearn/datasets/data/fsaverage5/sulc.left.gii.gz
9.2.16.3. Extracting the seed time series¶
# Load resting state time series from nilearn
from nilearn import surface
timeseries = surface.load_surf_data(nki_dataset['func_left'][0])
# Extract seed region via label
pcc_region = b'G_cingul-Post-dorsal'
import numpy as np
pcc_labels = np.where(parcellation == labels.index(pcc_region))[0]
# Extract time series from seed region
seed_timeseries = np.mean(timeseries[pcc_labels], axis=0)
9.2.16.4. Calculating seed-based functional connectivity¶
# Calculate Pearson product-moment correlation coefficient between seed
# time series and timeseries of all cortical nodes of the hemisphere
from scipy import stats
stat_map = np.zeros(timeseries.shape[0])
for i in range(timeseries.shape[0]):
stat_map[i] = stats.pearsonr(seed_timeseries, timeseries[i])[0]
# Re-mask previously masked nodes (medial wall)
stat_map[np.where(np.mean(timeseries, axis=1) == 0)] = 0
Out:
/usr/lib/python3/dist-packages/scipy/stats/stats.py:3399: PearsonRConstantInputWarning: An input array is constant; the correlation coefficent is not defined.
warnings.warn(PearsonRConstantInputWarning())
Display ROI on surface
# Transform ROI indices in ROI map
pcc_map = np.zeros(parcellation.shape[0], dtype=int)
pcc_map[pcc_labels] = 1
from nilearn import plotting
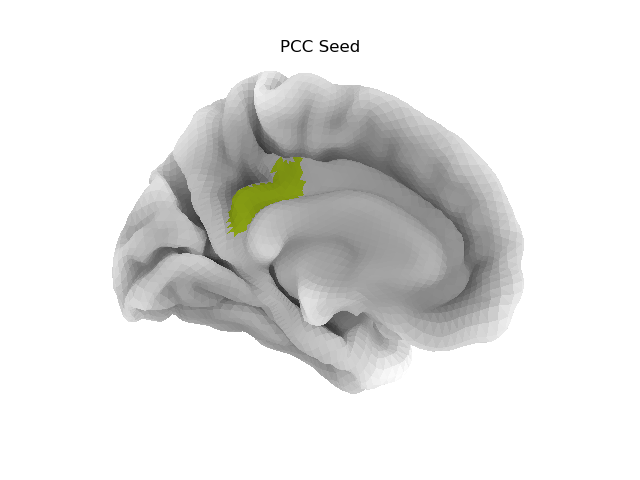
plotting.plot_surf_roi(fsaverage['pial_left'], roi_map=pcc_map,
hemi='left', view='medial',
bg_map=fsaverage['sulc_left'], bg_on_data=True,
title='PCC Seed')
Out:
<Figure size 640x480 with 1 Axes>
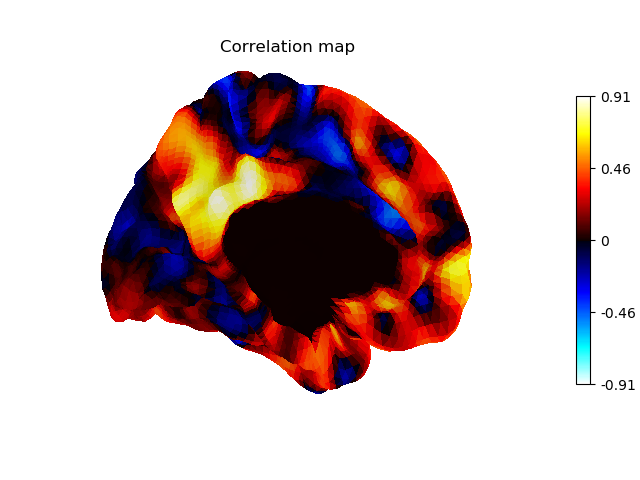
Display unthresholded stat map with a slightly dimmed background
plotting.plot_surf_stat_map(fsaverage['pial_left'], stat_map=stat_map,
hemi='left', view='medial', colorbar=True,
bg_map=fsaverage['sulc_left'], bg_on_data=True,
darkness=.3, title='Correlation map')
Out:
<Figure size 640x480 with 2 Axes>
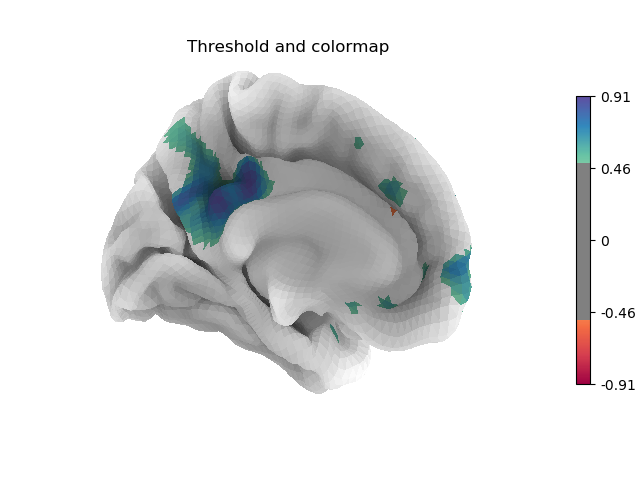
Many different options are available for plotting, for example thresholding, or using custom colormaps
plotting.plot_surf_stat_map(fsaverage['pial_left'], stat_map=stat_map,
hemi='left', view='medial', colorbar=True,
bg_map=fsaverage['sulc_left'], bg_on_data=True,
cmap='Spectral', threshold=.5,
title='Threshold and colormap')
Out:
<Figure size 640x480 with 2 Axes>
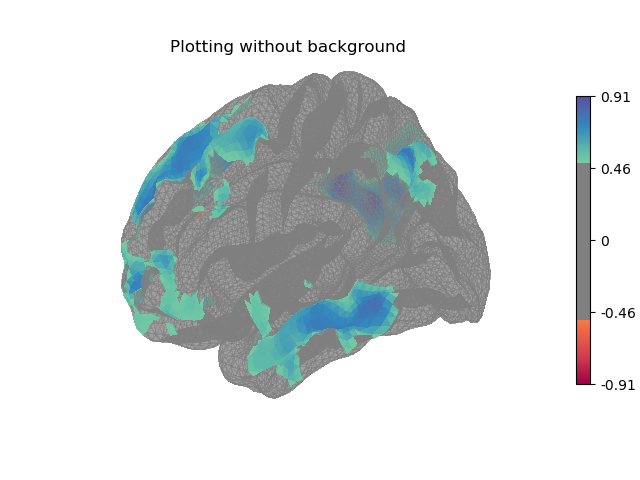
Here the surface is plotted in a lateral view without a background map. To capture 3D structure without depth information, the default is to plot a half transparent surface. Note that you can also control the transparency with a background map using the alpha parameter.
plotting.plot_surf_stat_map(fsaverage['pial_left'], stat_map=stat_map,
hemi='left', view='lateral', colorbar=True,
cmap='Spectral', threshold=.5,
title='Plotting without background')
Out:
<Figure size 640x480 with 2 Axes>
The plots can be saved to file, in which case the display is closed after creating the figure
plotting.plot_surf_stat_map(fsaverage['infl_left'], stat_map=stat_map,
hemi='left', bg_map=fsaverage['sulc_left'],
bg_on_data=True, threshold=.5, colorbar=True,
output_file='plot_surf_stat_map.png')
plotting.show()
Total running time of the script: ( 1 minutes 5.328 seconds)
